Diff Studio provides a simulation of multi-stage processes. Due to the new Diff Grok features, parameter optimization for such models runs in parallel now:
Diff Studio enables modeling cyclic processes, in particular pharmacokinetic-pharmacodynamics. The new version of the Diff Grok library provides parallel parameters optimization for such models.
Check the new release of Diff Studio.
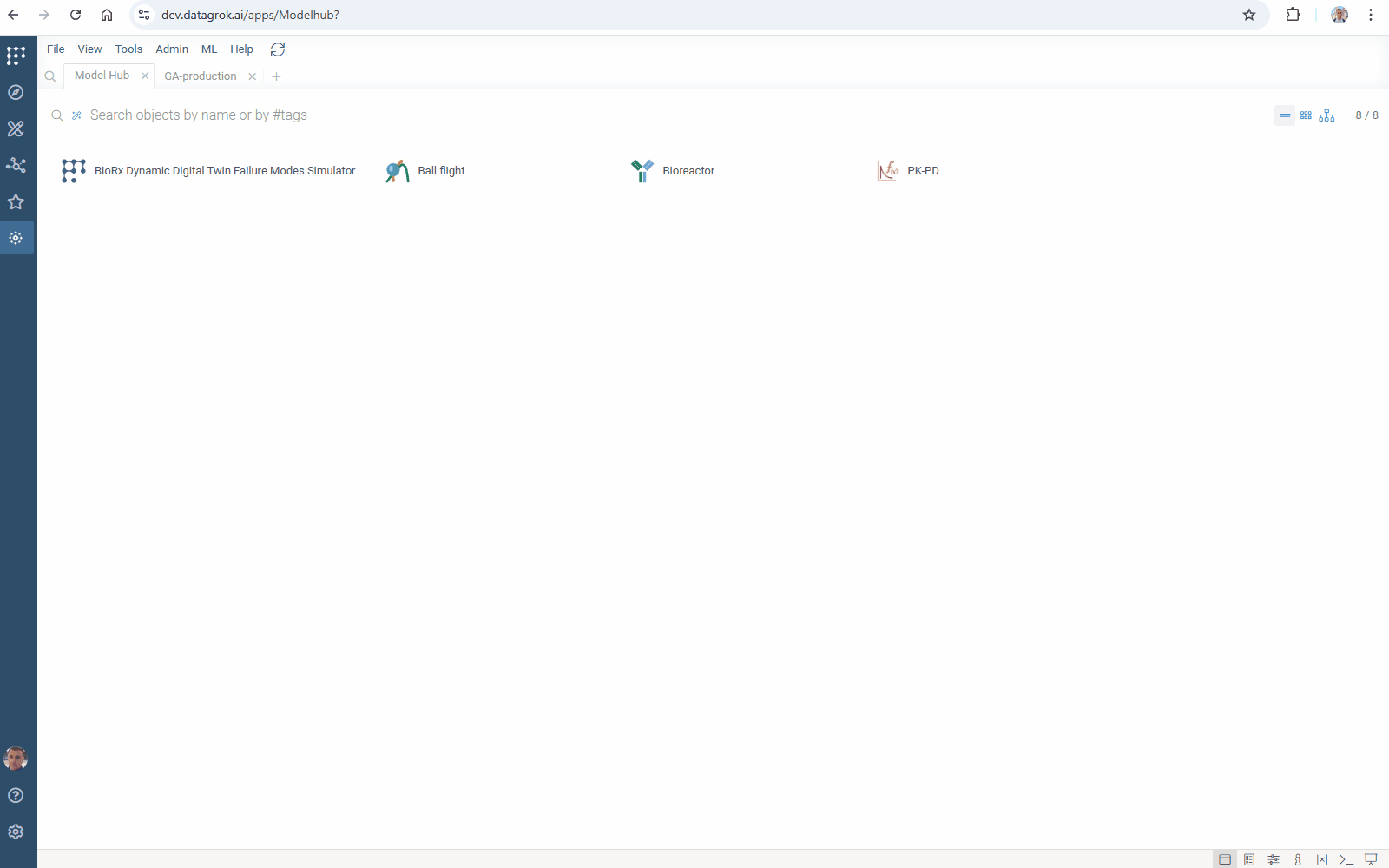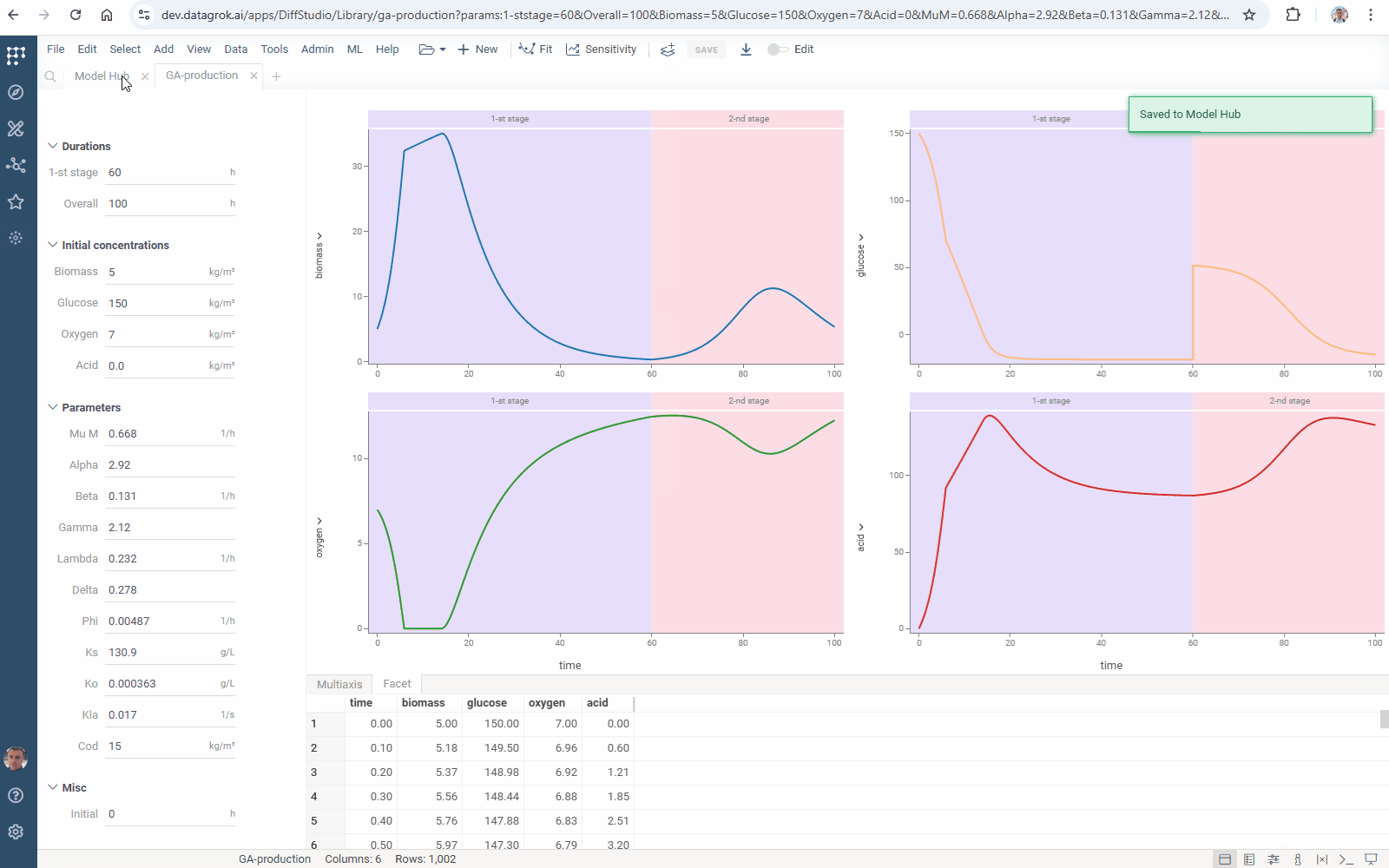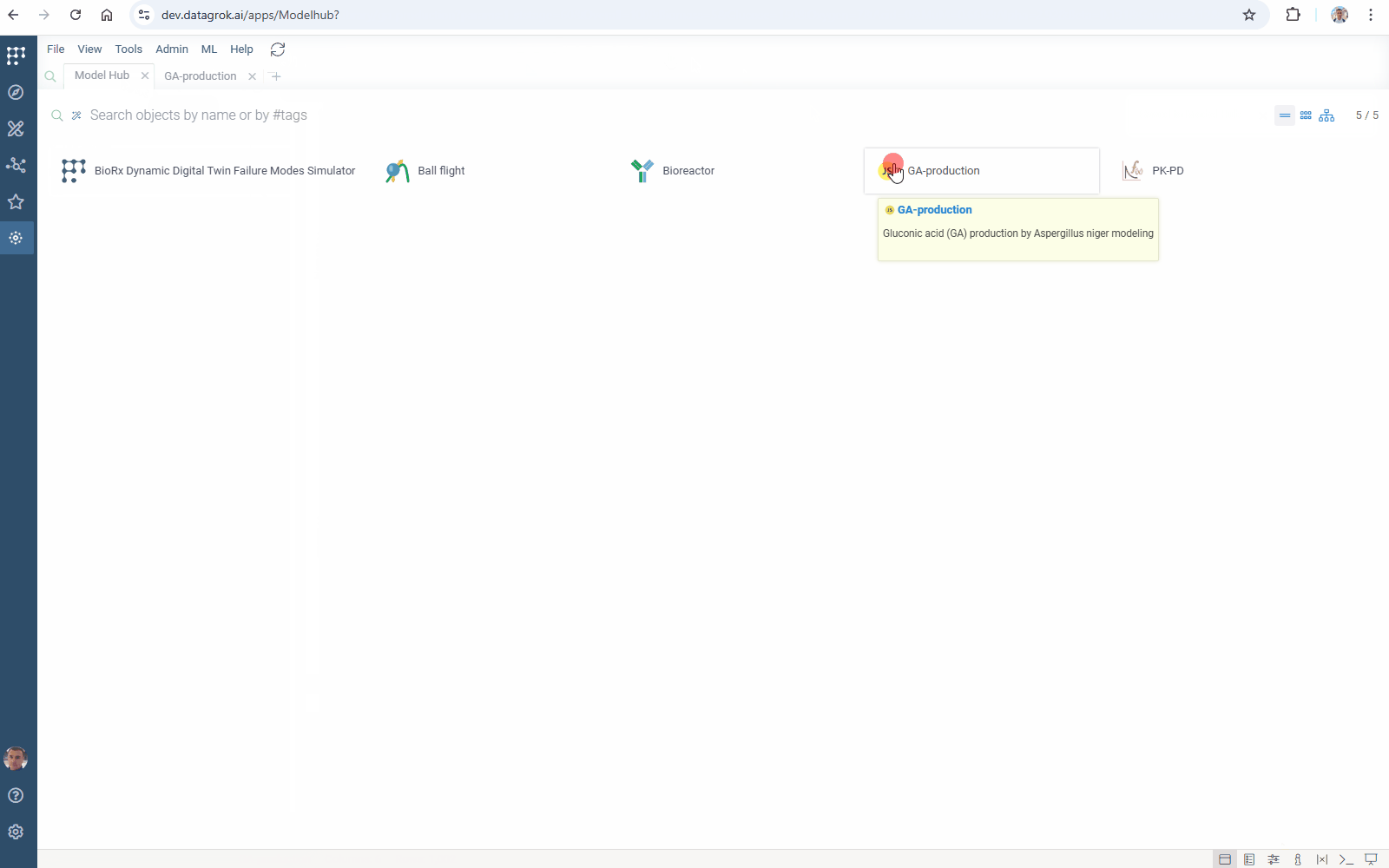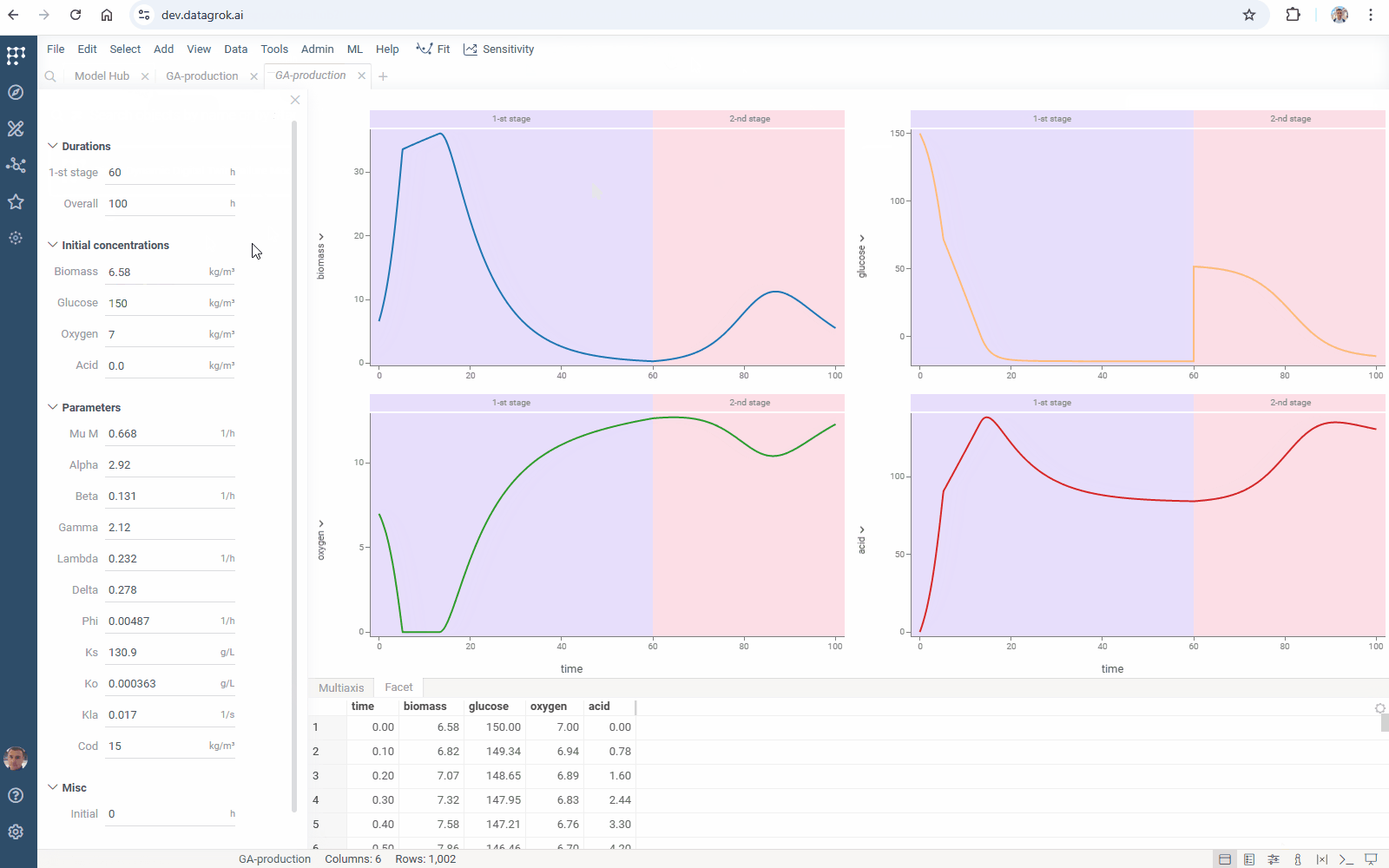
You can add your model to Model Hub in one click:
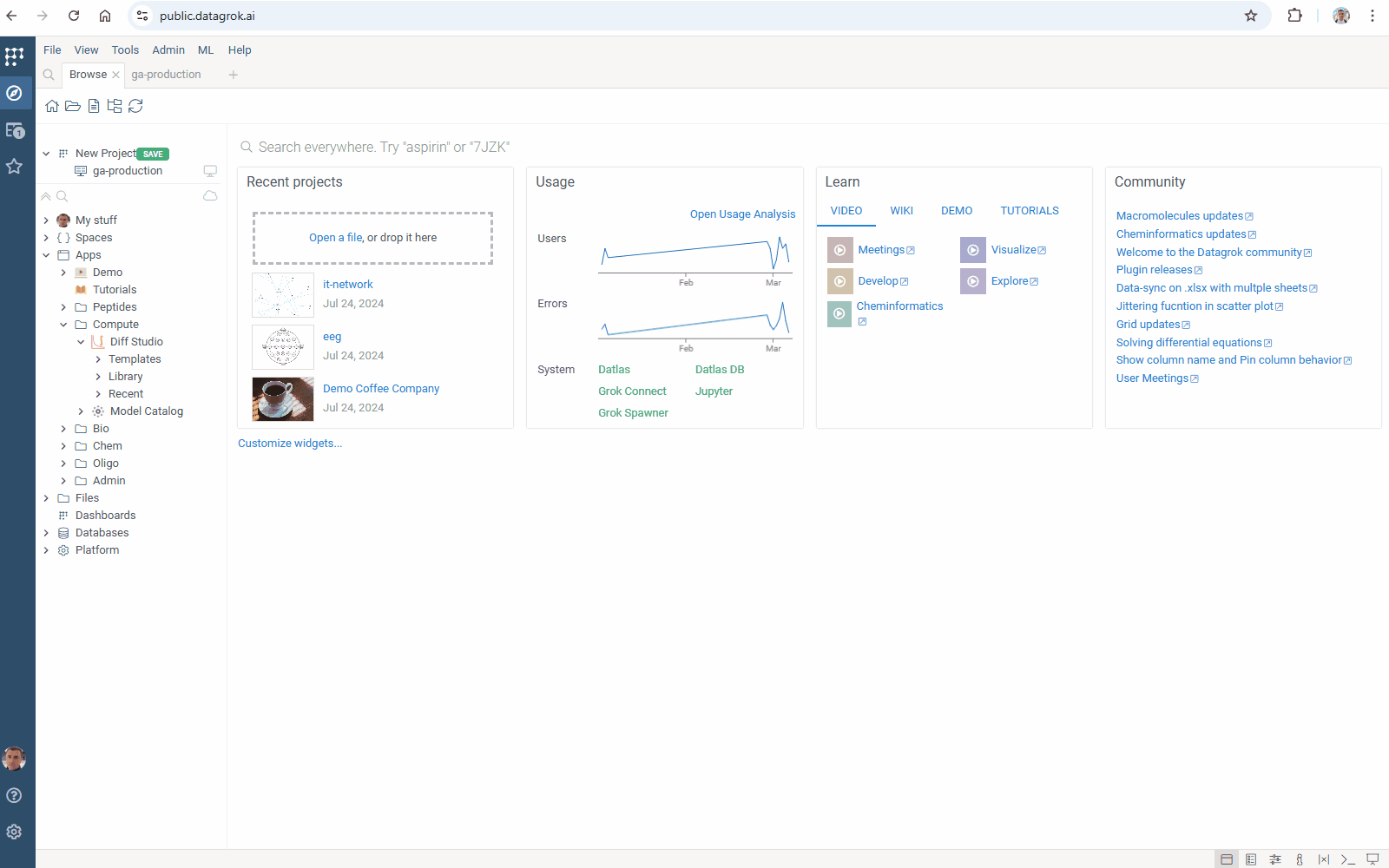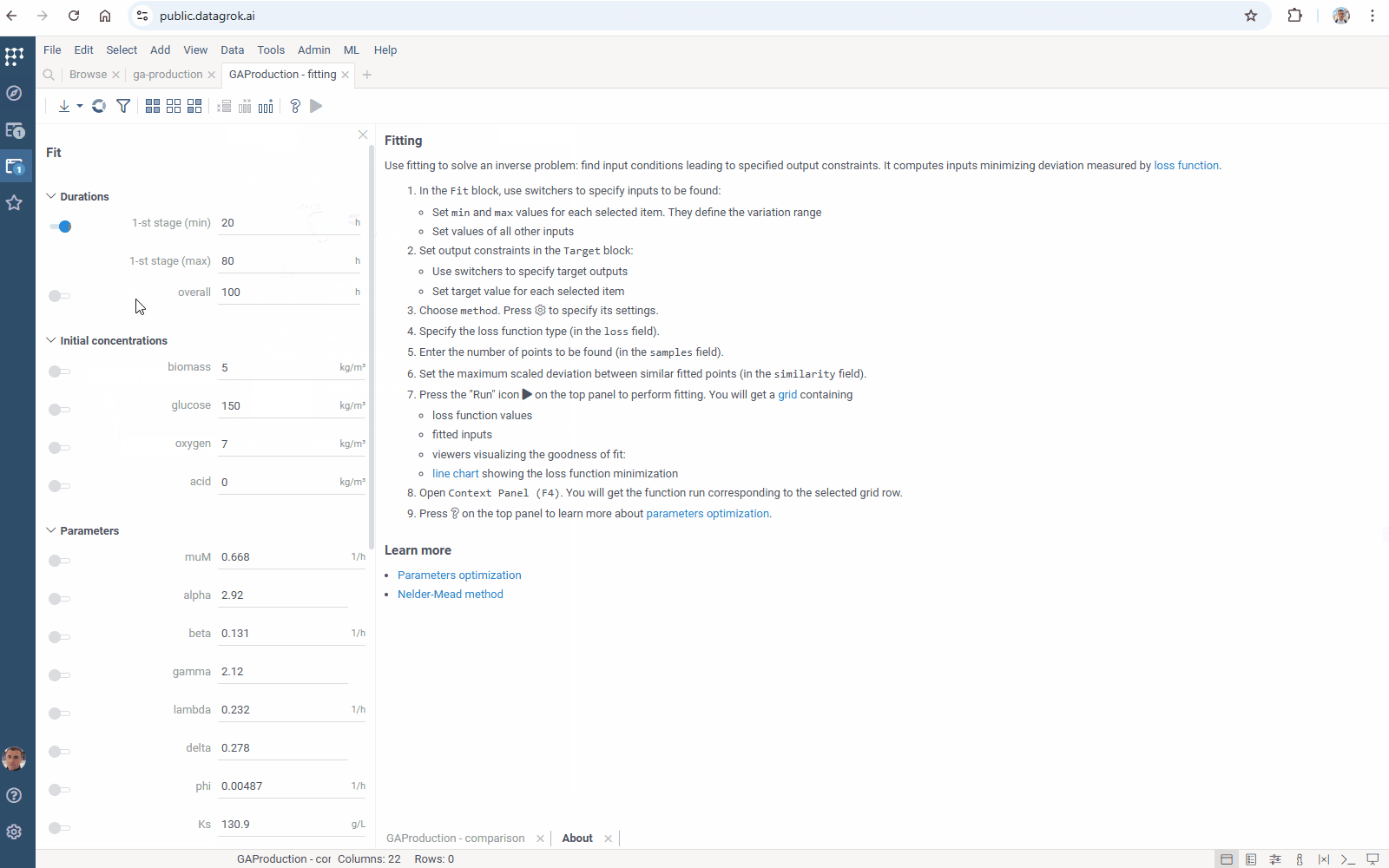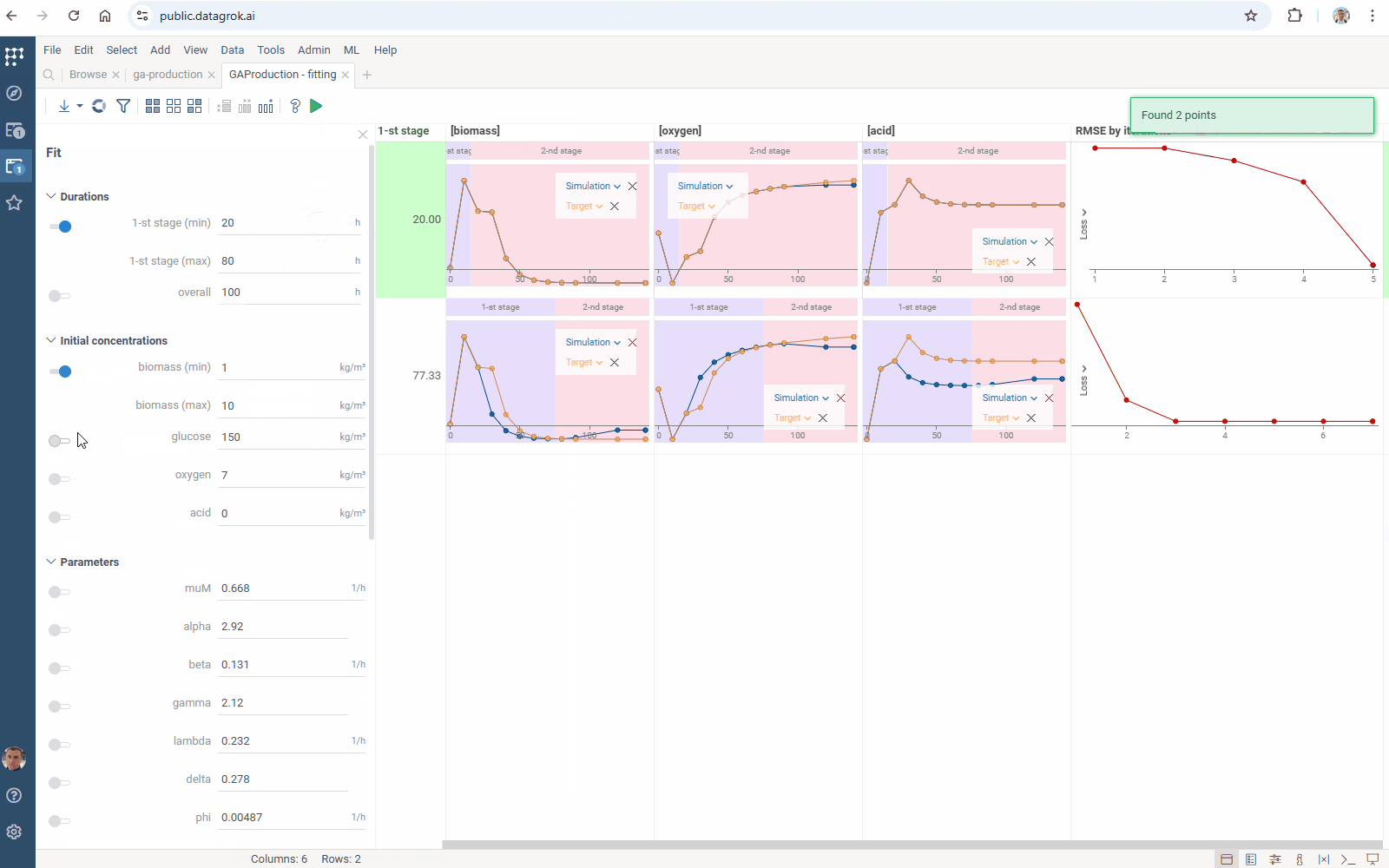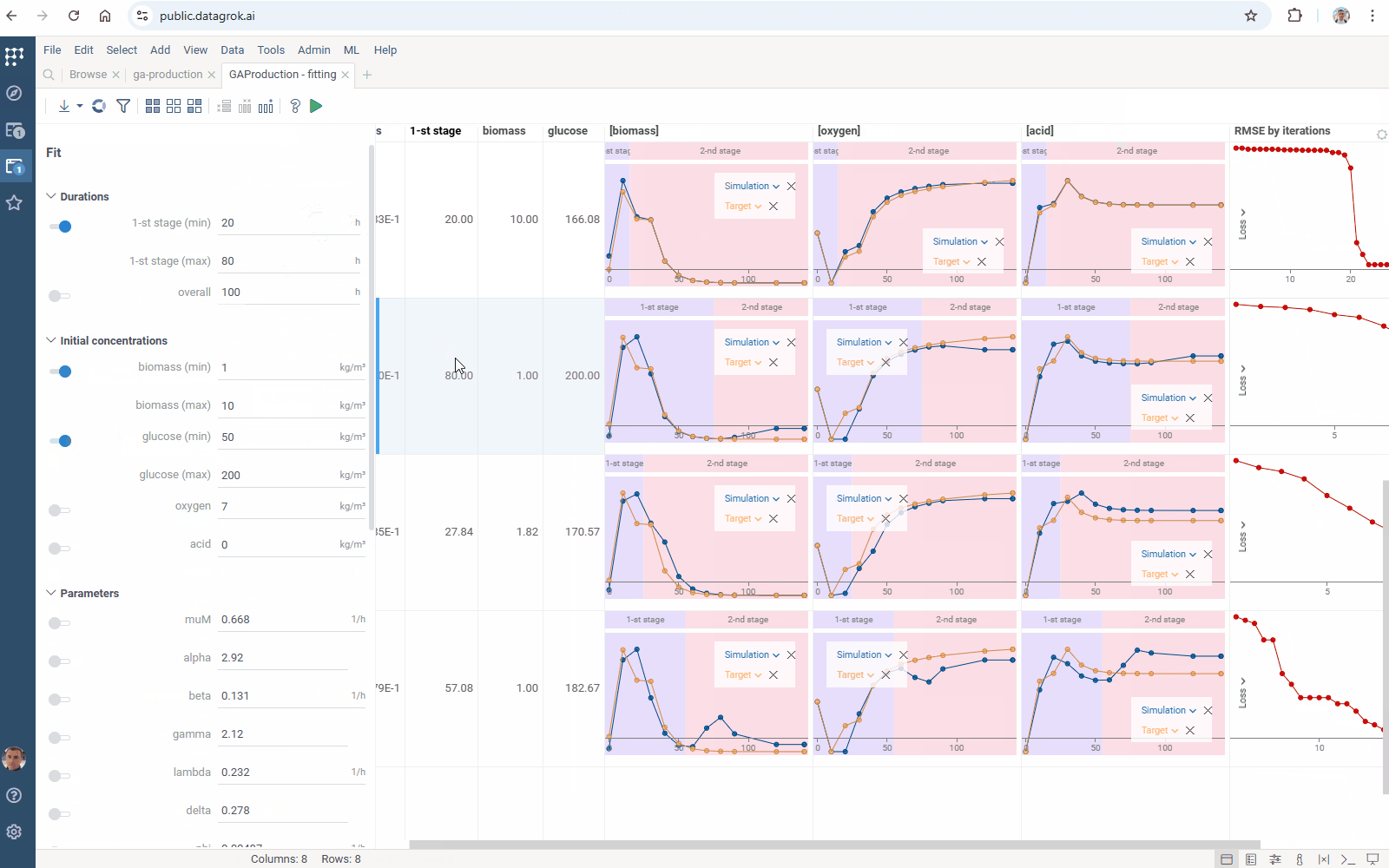
Meet improved parameters optimization for Diff Studio models:
- Flexibility: select the target function columns of your table with experimental data
- Output management: filter similar results to get the most different curves
- Usability: use navigation when specifying the target output
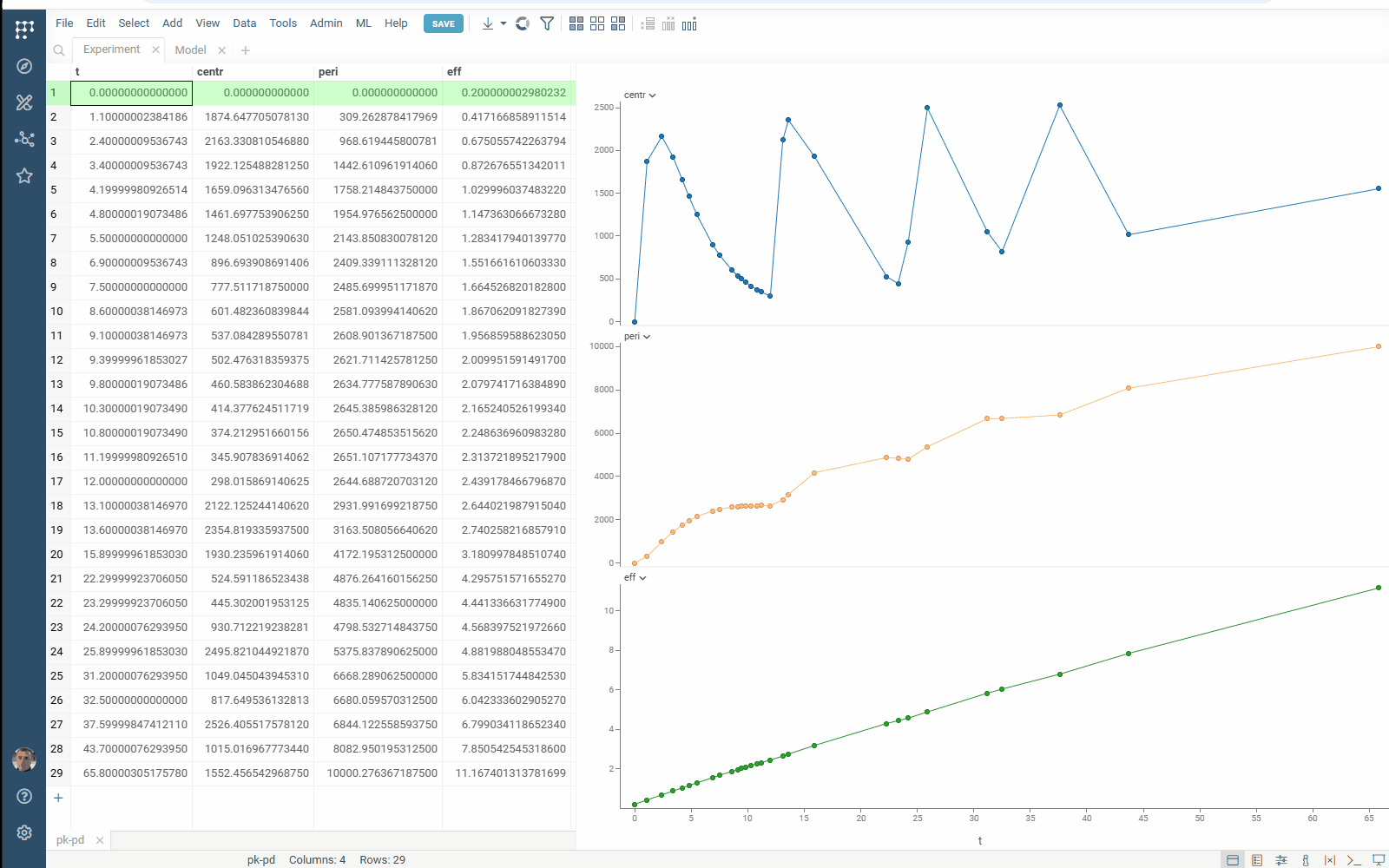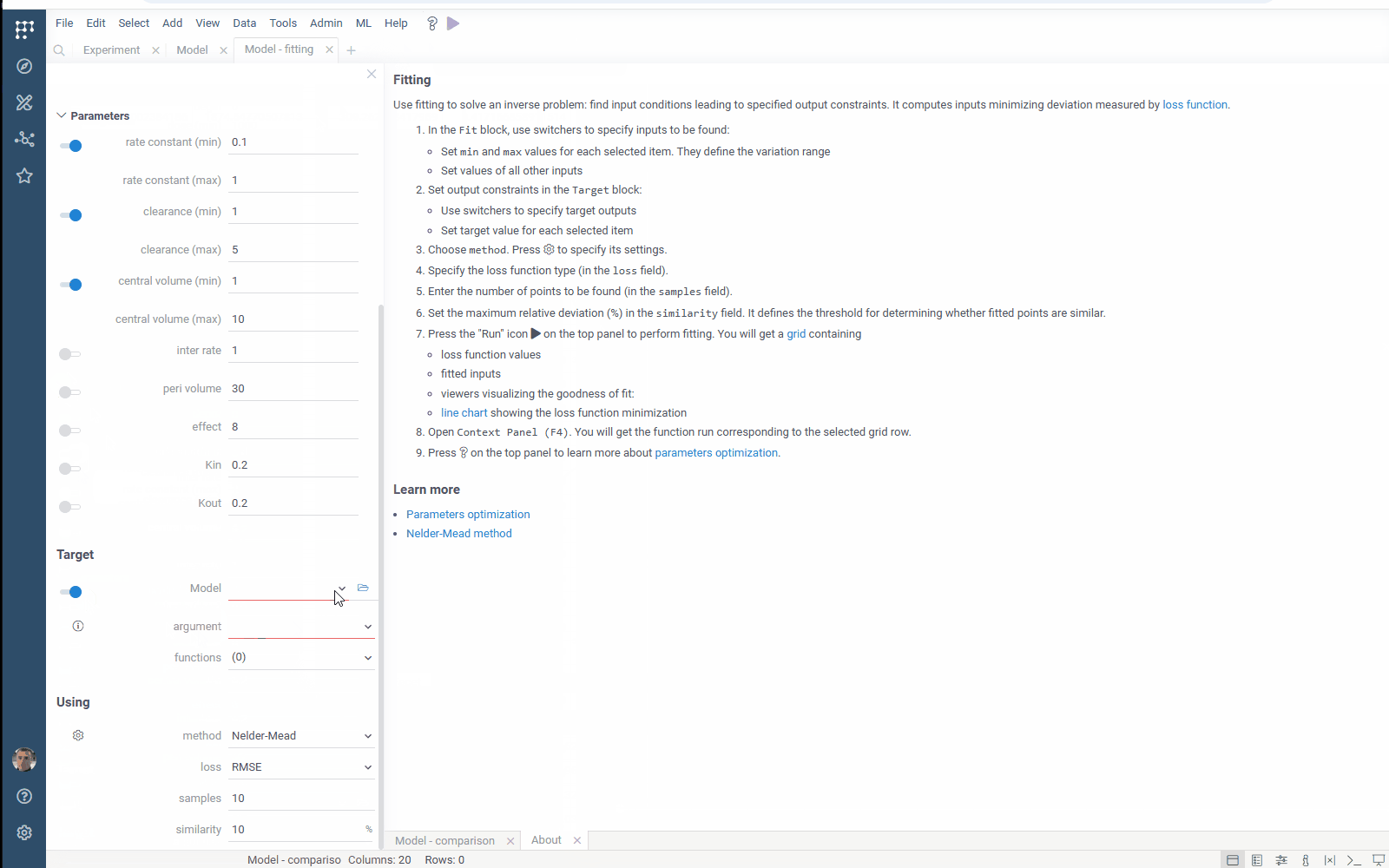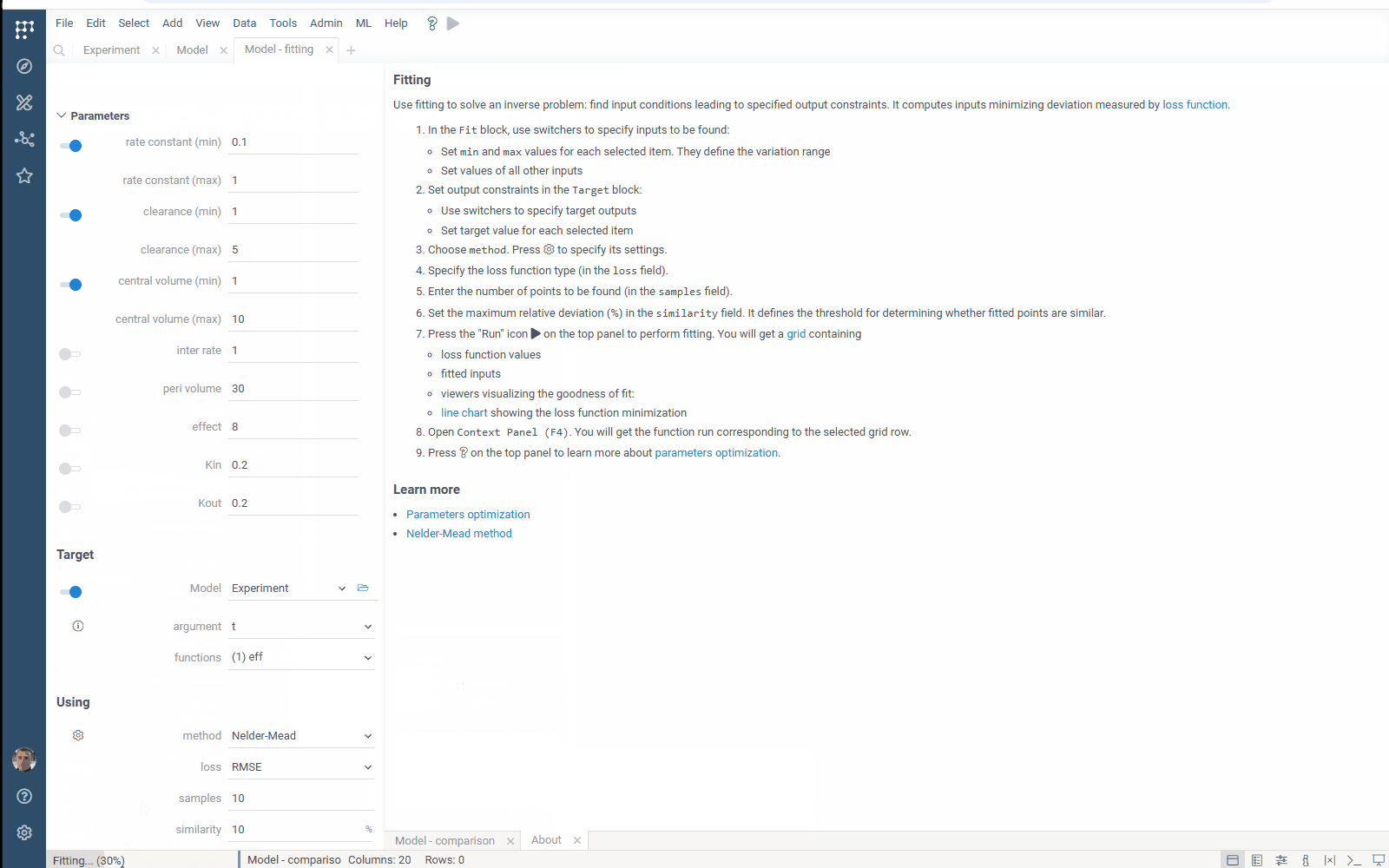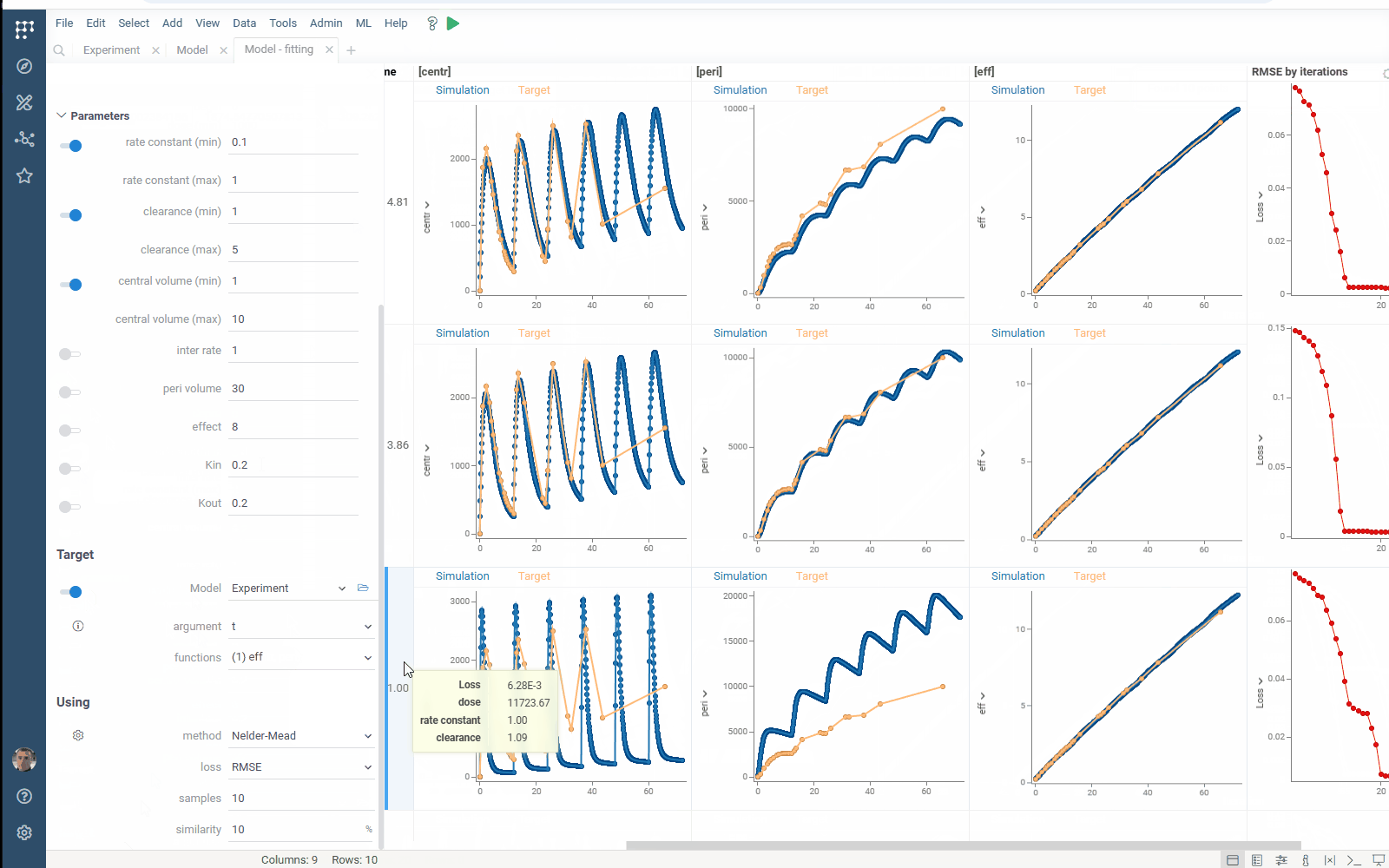
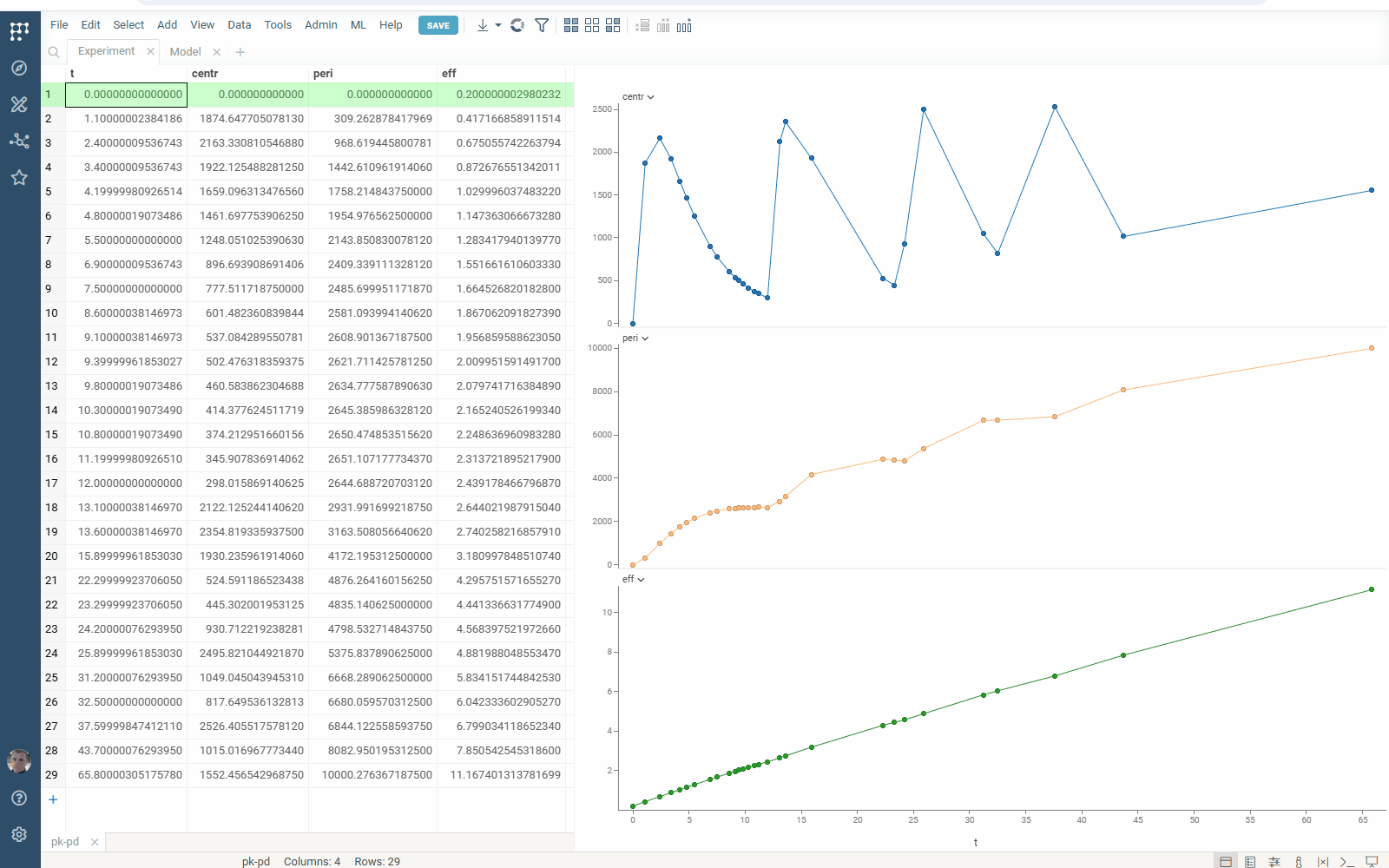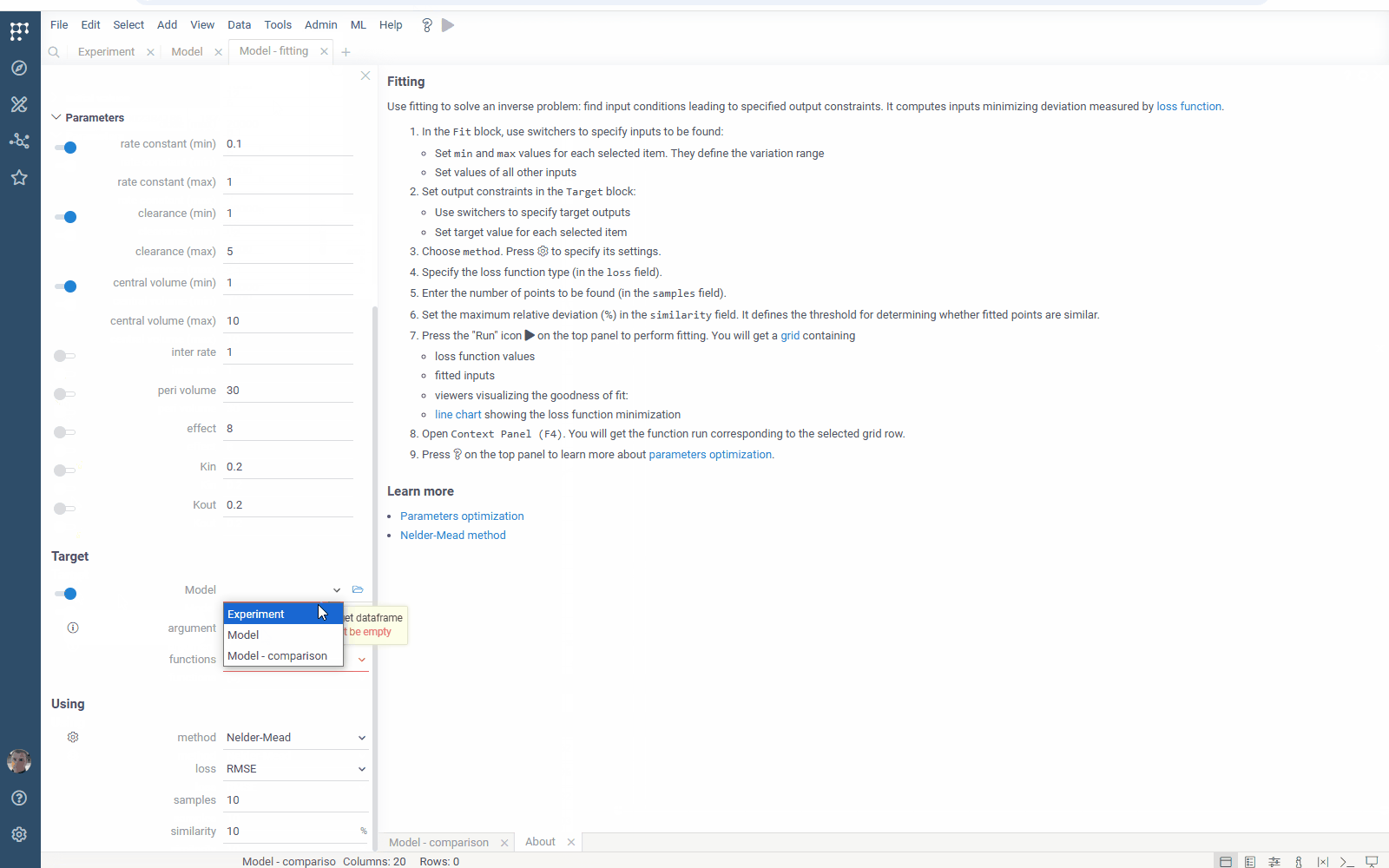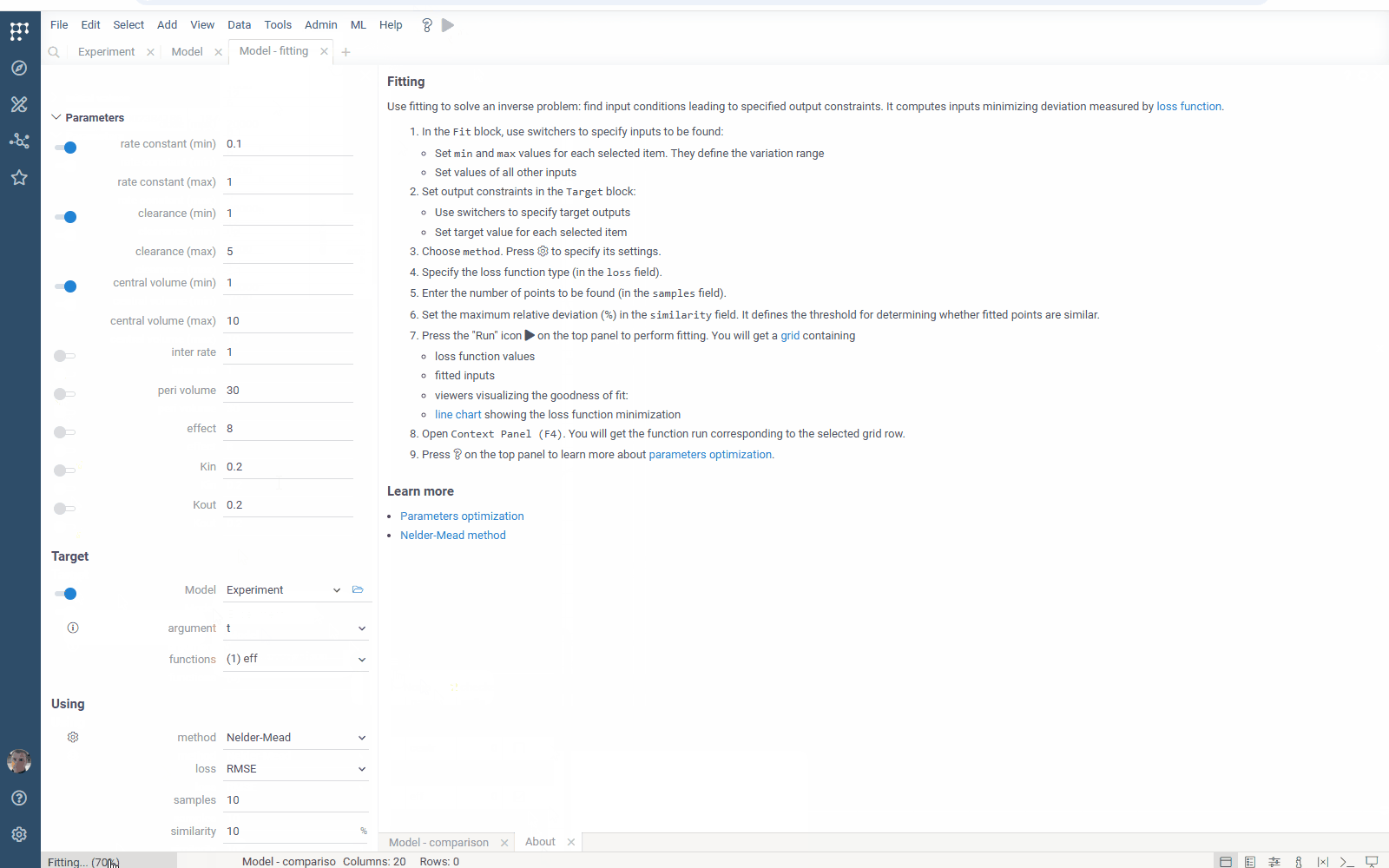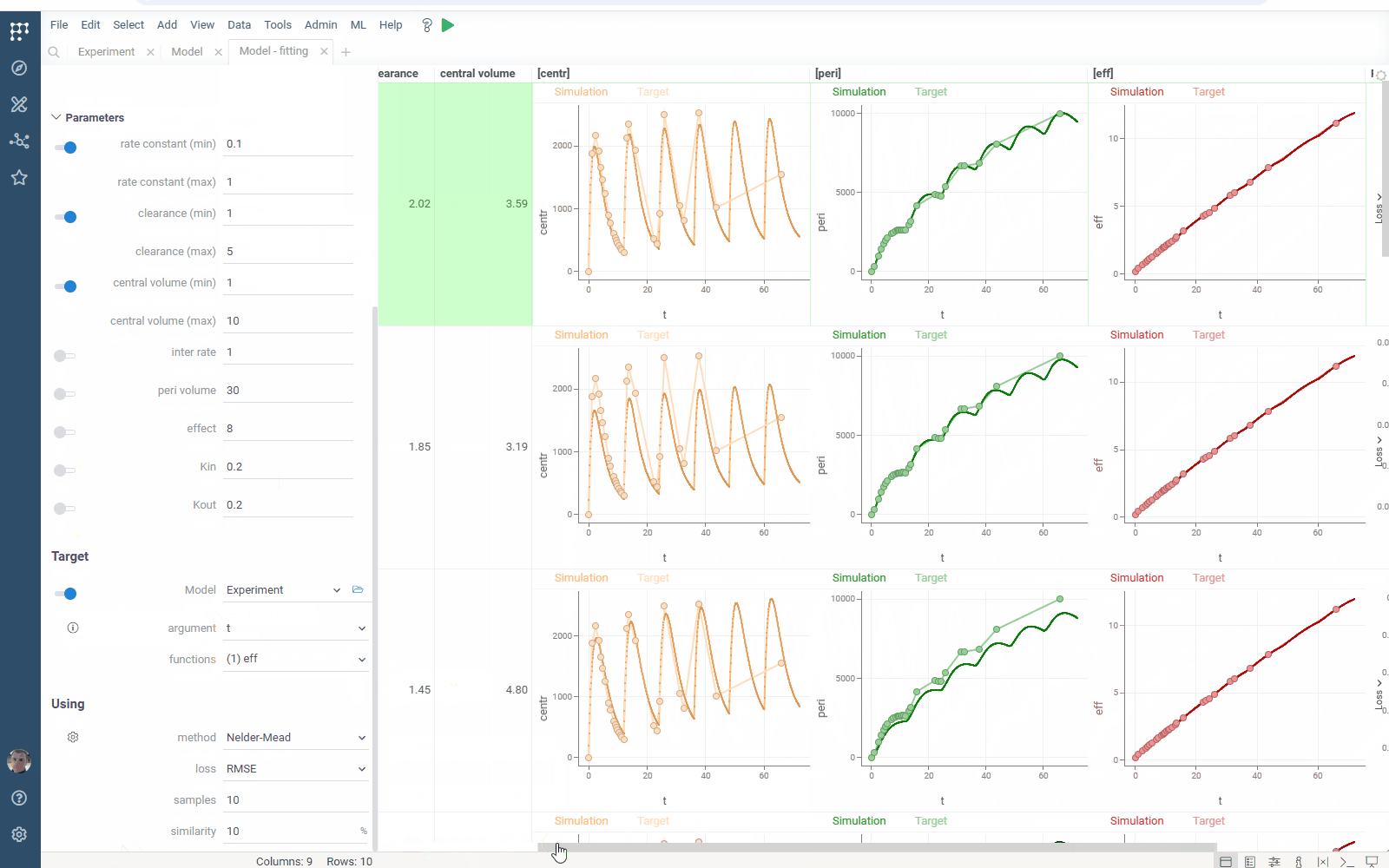
Check the improved model analysis. Explore results of parameter optimization in a more convenient way:
Meet accelerated sensitivity analysis for Diff Studio models. Use viewers’ features to explore the impact of model input variation on its output:
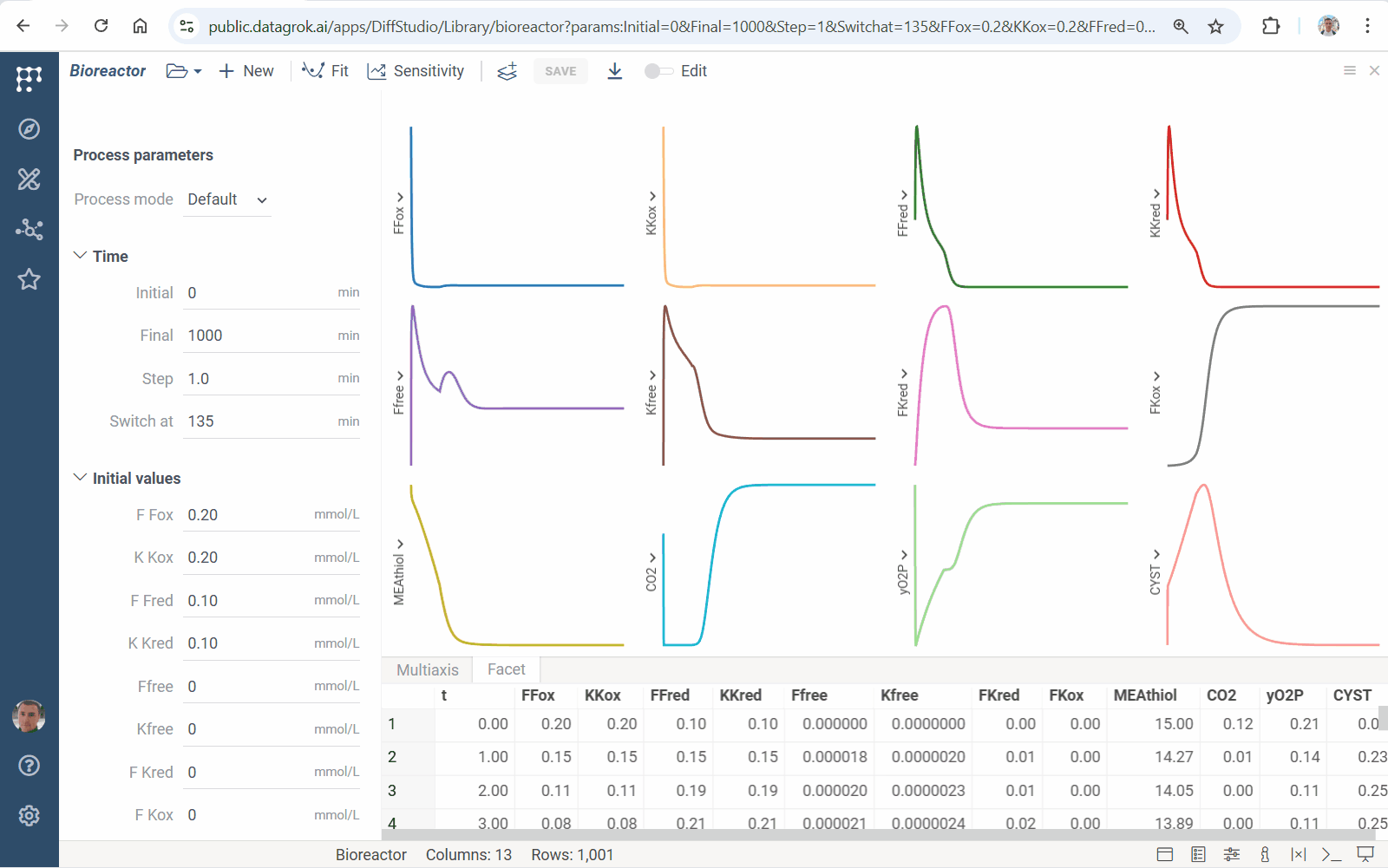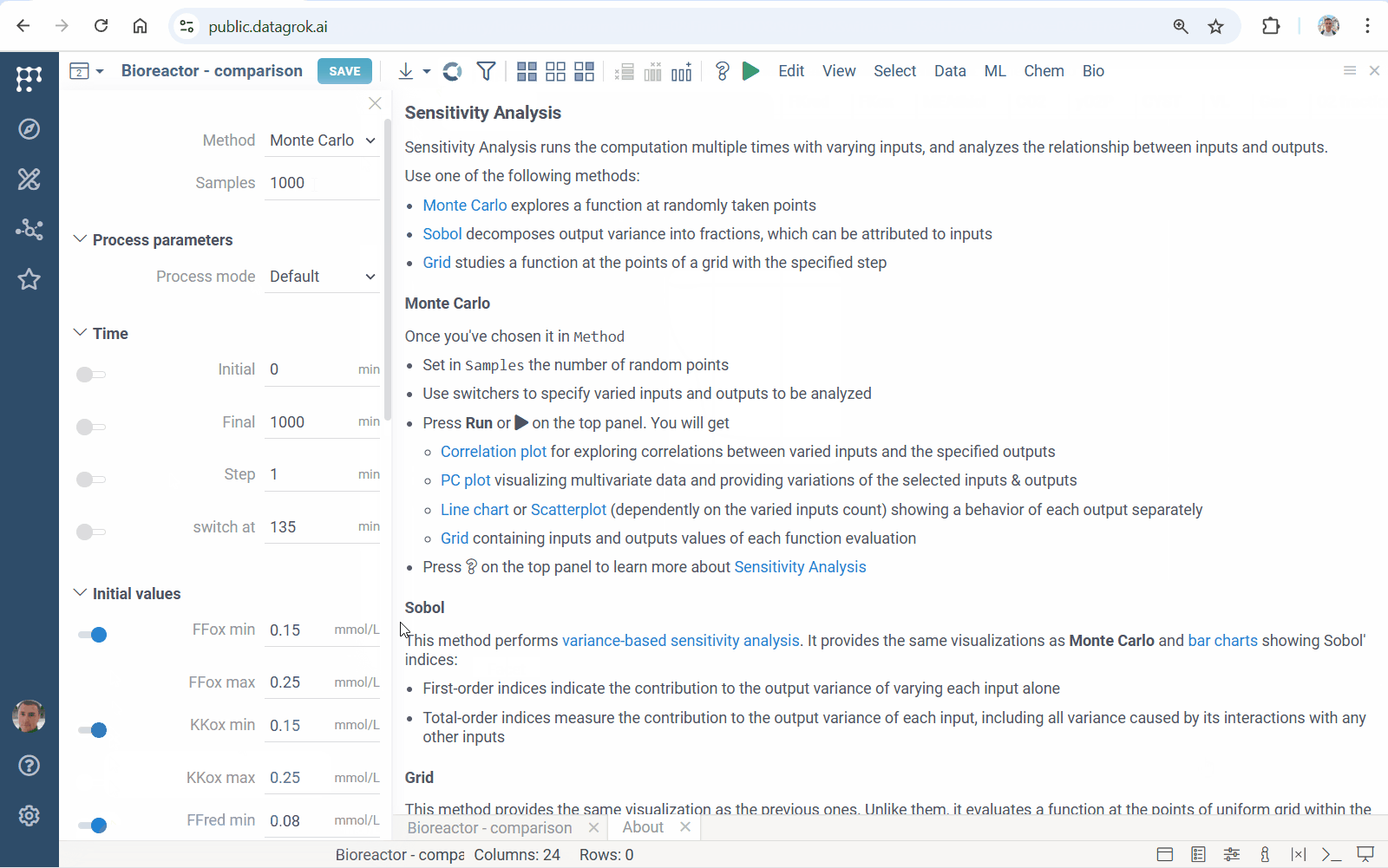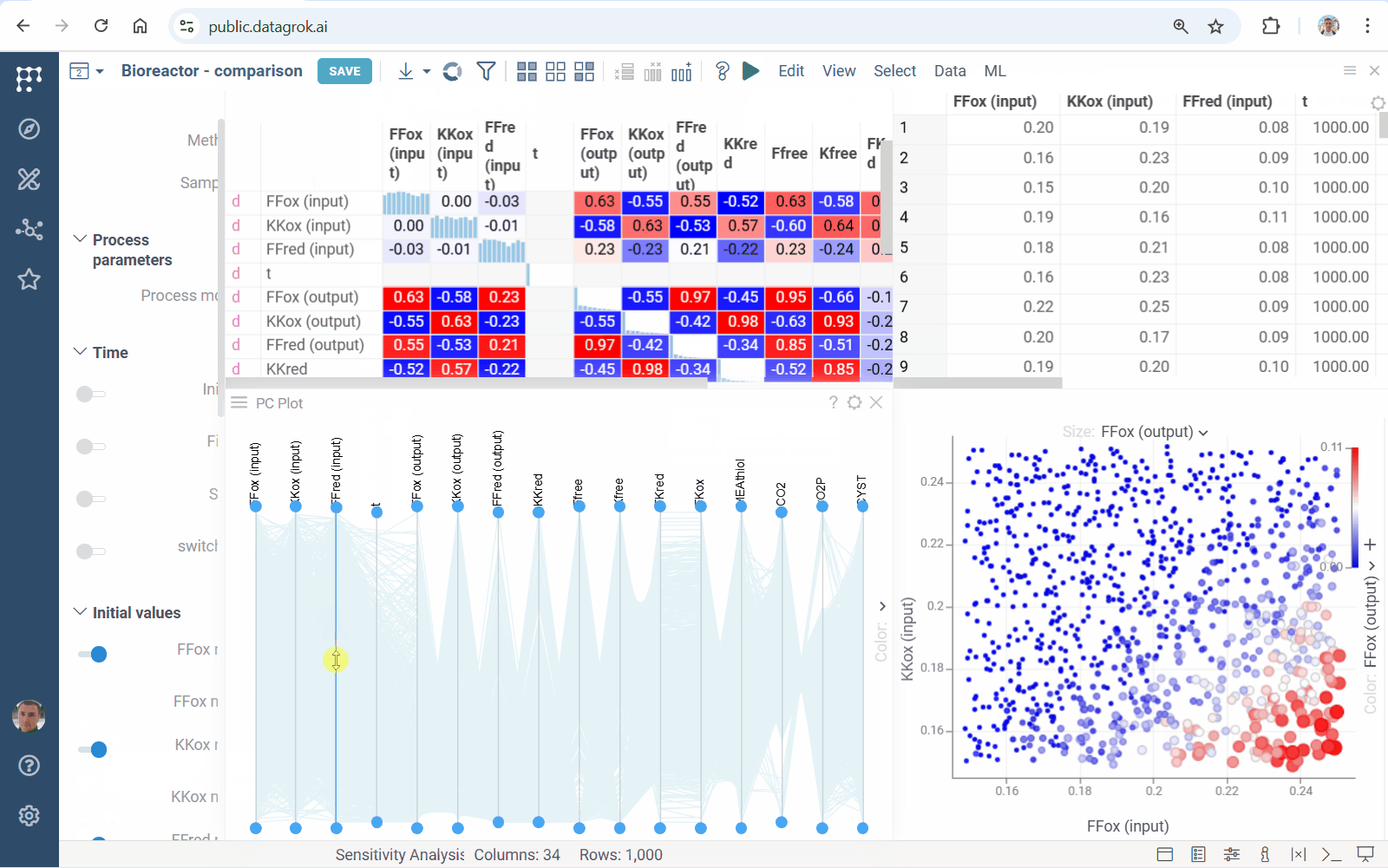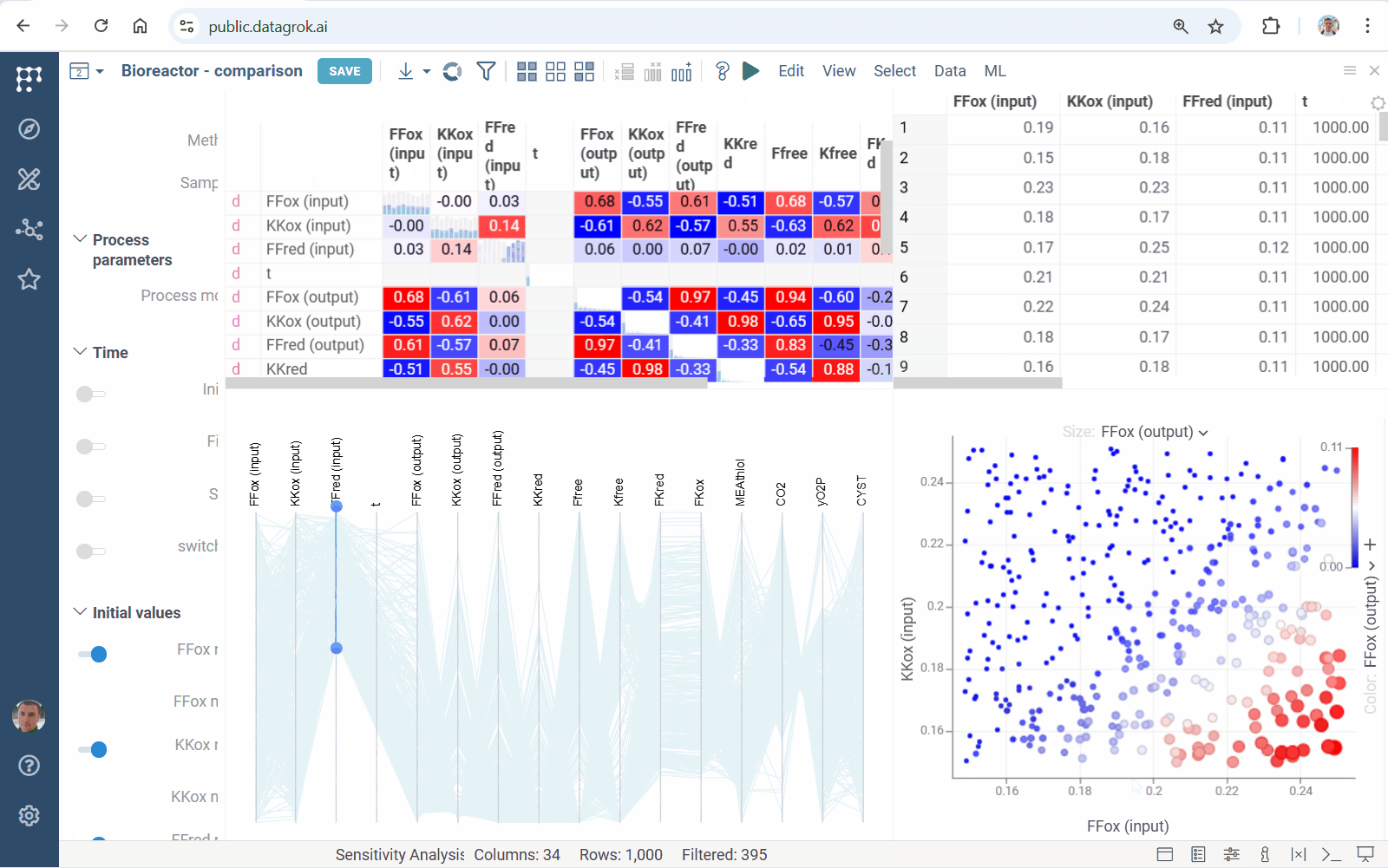
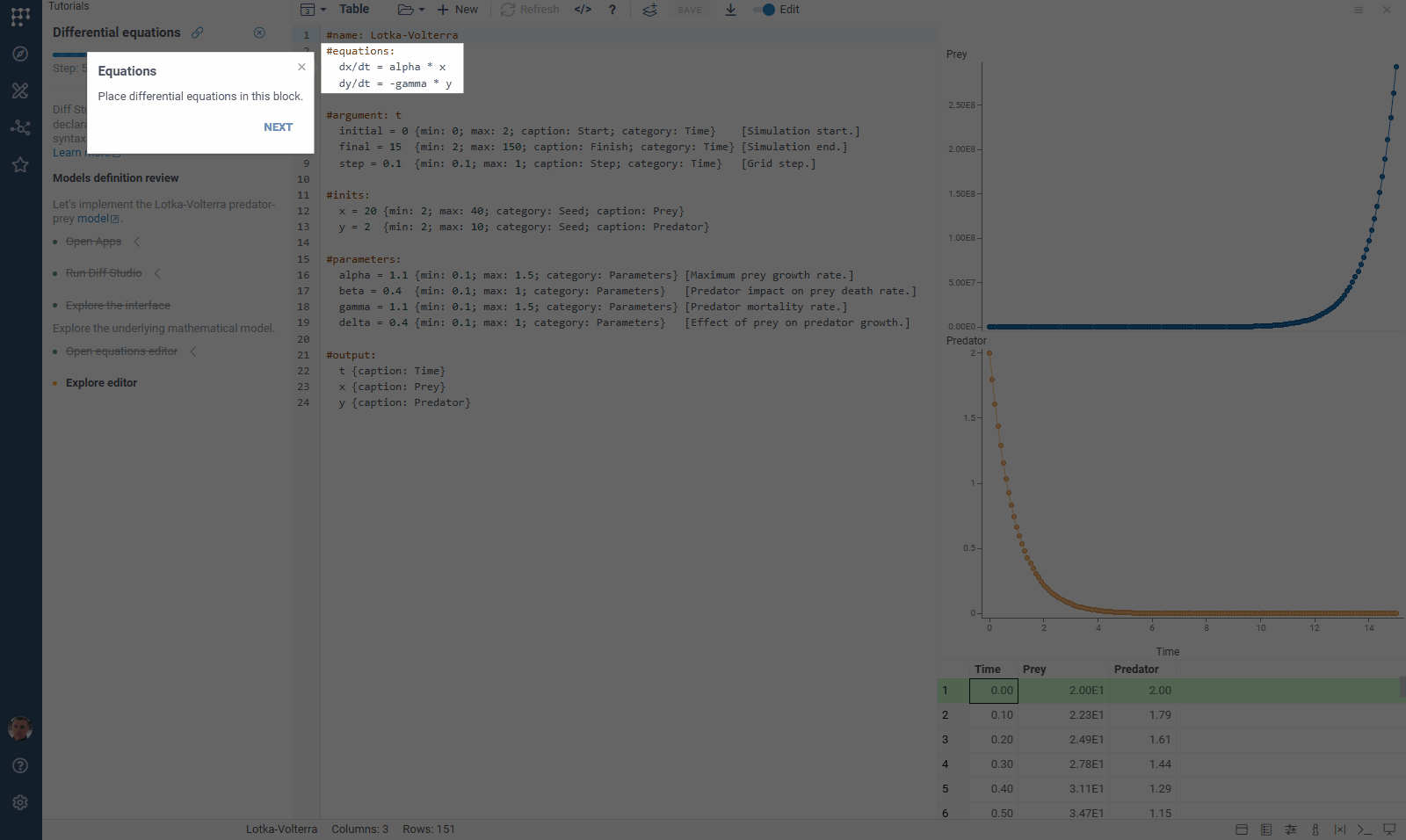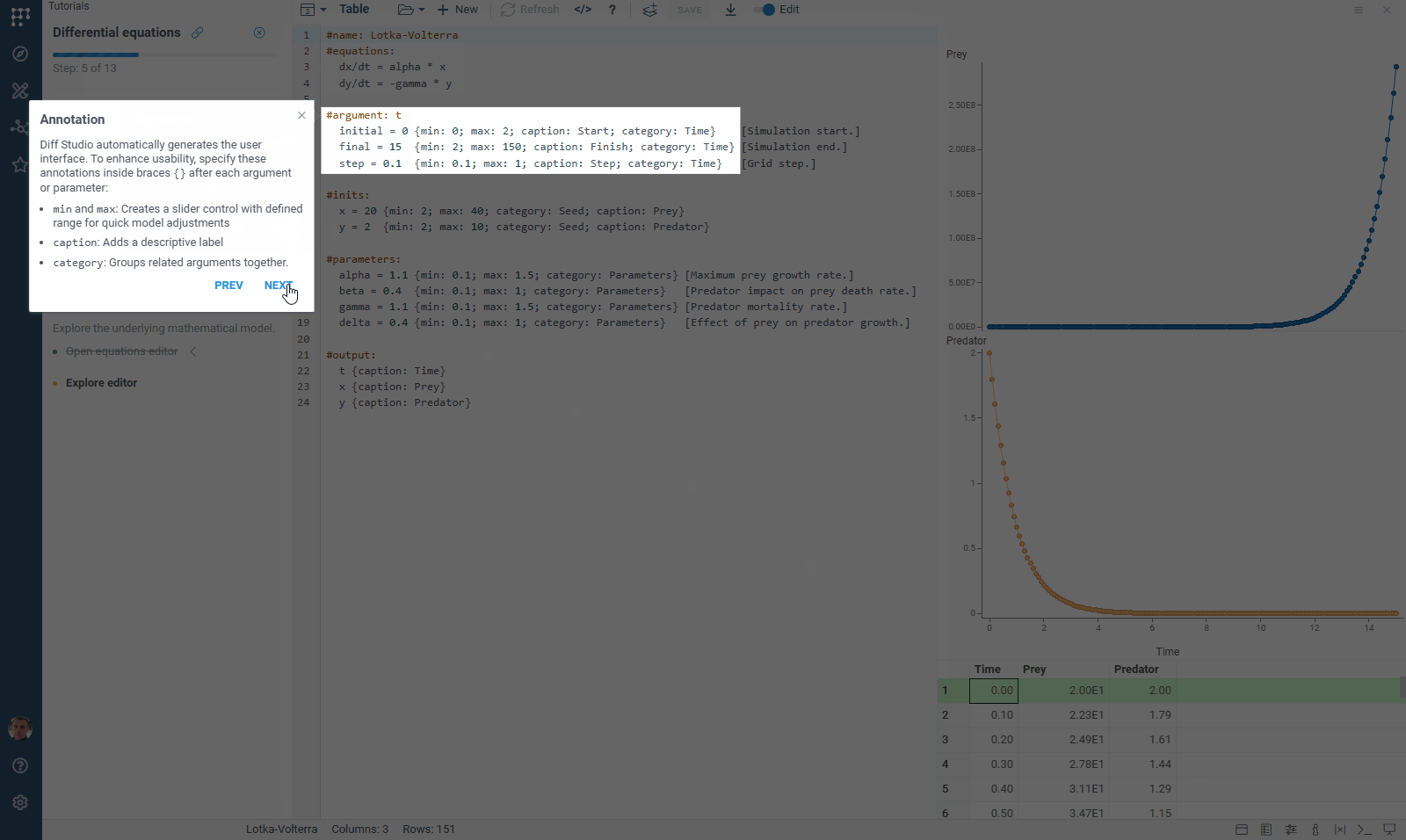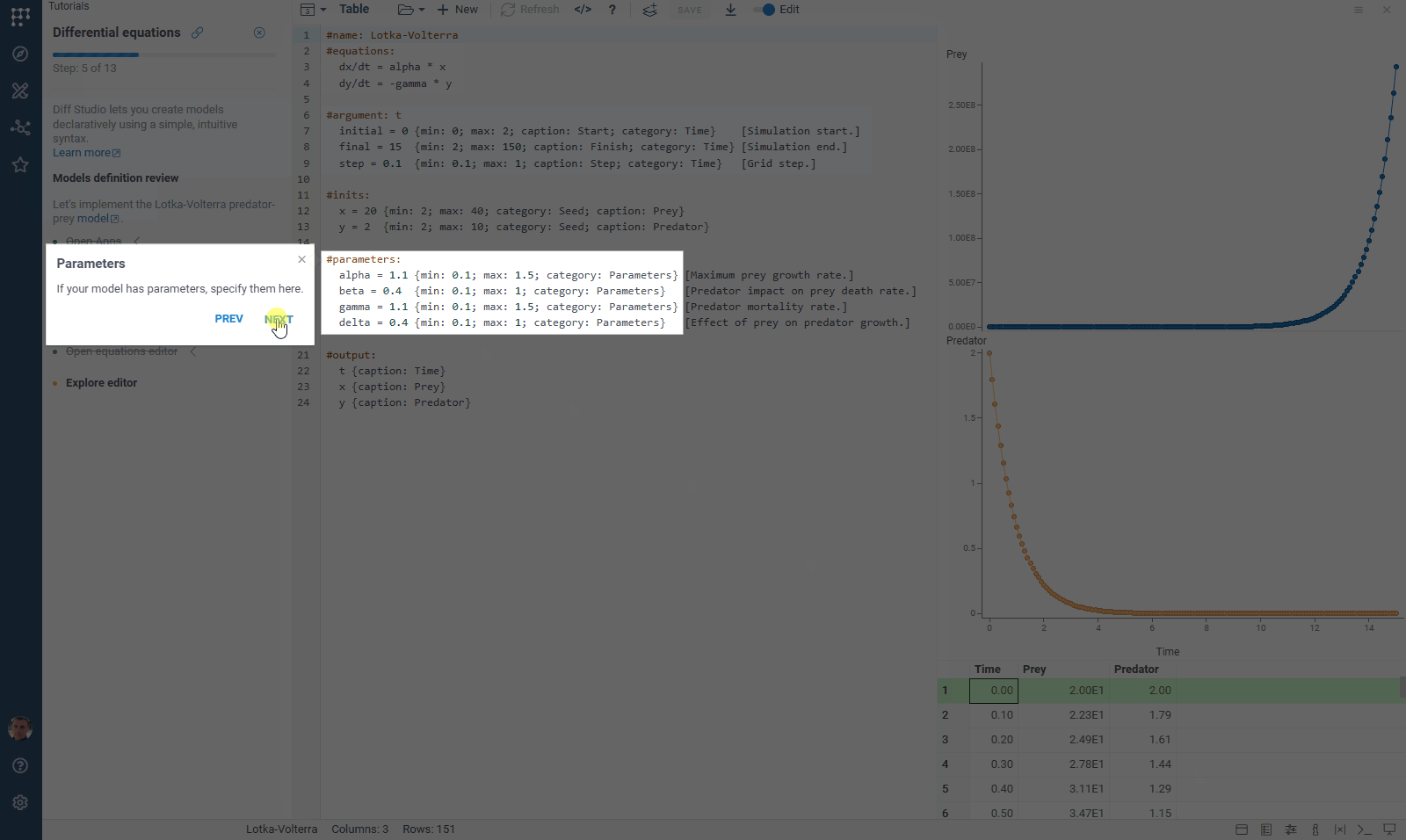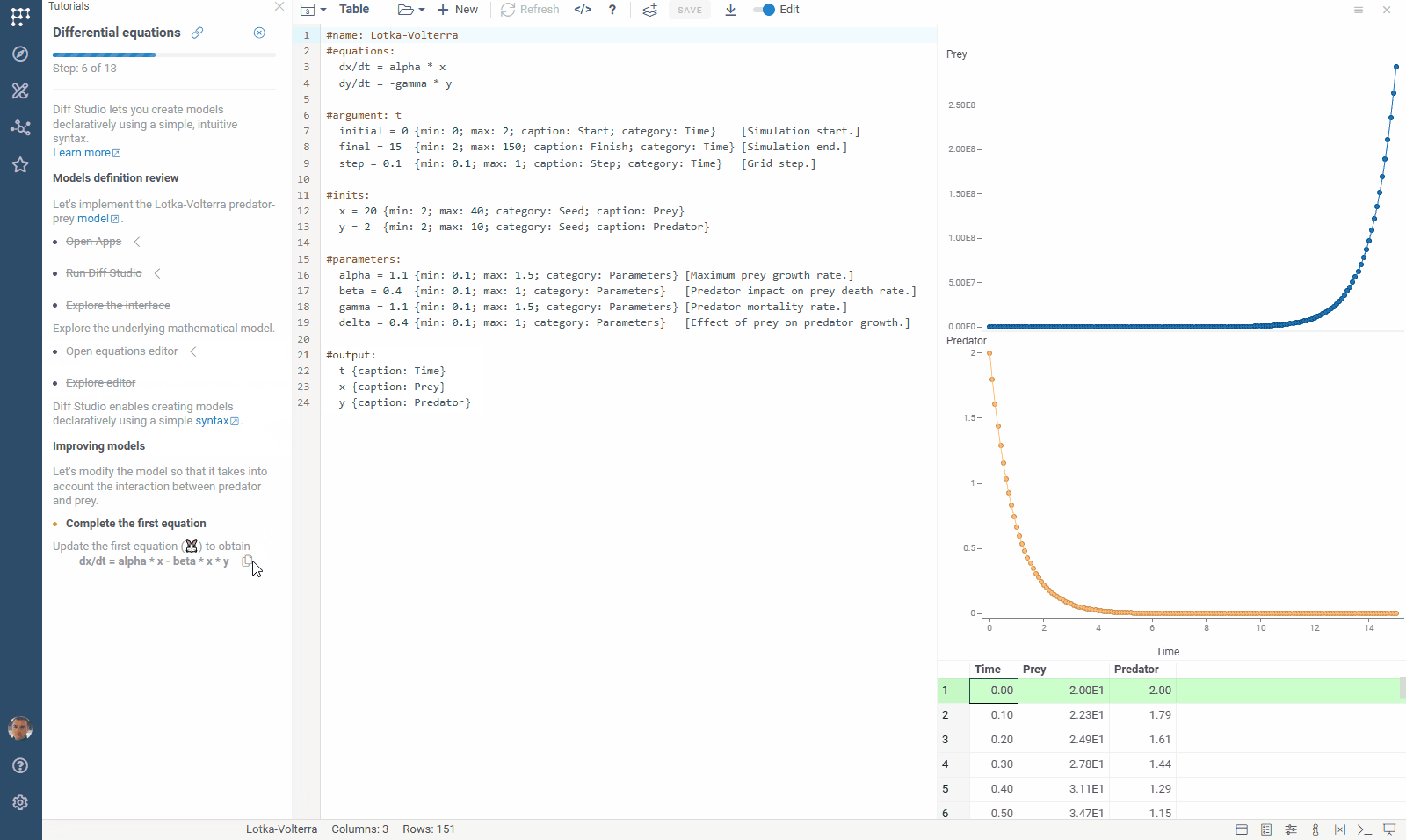
We’ve updated our Diff Studio tutorial! Check it out here. The new version improves navigation through model components:
Diff Grok now supports 9 numerical methods for solving ODEs in the browser. Six new methods join the existing Rosenbrock-Wanner solvers:
- Runge-Kutta - for non-stiff ODEs:
- Bogacki-Shampine 3(2)
- Runge-Kutta-Fehlberg 4(5)
- Dormand-Prince 5(4)
- Adams - multistep predictor-correctors of order 4 and 5
- LSODA - automatic stiffness detection
Diff Studio now supports 10 numerical methods for solving ODEs in the browser. Seven new methods join the existing Rosenbrock-Wanner solvers:
- Runge-Kutta & Adams - for non-stiff ODEs: Bogacki-Shampine 3(2), Runge-Kutta-Fehlberg 4(5), Dormand-Prince 5(4), and multistep predictor-correctors of order 4 and 5
- LSODA, CVODE - adaptive stiff/non-stiff solvers
Match the solver to your problem: Runge-Kutta for smooth systems, Adams when function evaluations are expensive, LSODA & CVODE when stiffness is unknown, and Rosenbrock-Wanner for known stiff equations.
Check it: Diff Studio.
Meet the new release of Diff Grok. Version 1.3.0 provides model export to publication-ready TeX and Markdown! ![]() You can now convert your ODE-based models directly into formats ready to paste into a paper, docs, or wiki.
You can now convert your ODE-based models directly into formats ready to paste into a paper, docs, or wiki.