CDD Vault Link 1.0.4 (2025-05-28) docs, release notes
Features:
- Implemented routing functionality
- Added ‘Protocols’ and ‘Collections’ links to the home page
Check out our new demo video to explore the full capabilities of the CDD Vault Link plugin.
3 Likes
Bio 2.22.0 ( 2025-06-03) docs, release notes
MSA Header Enhancements:
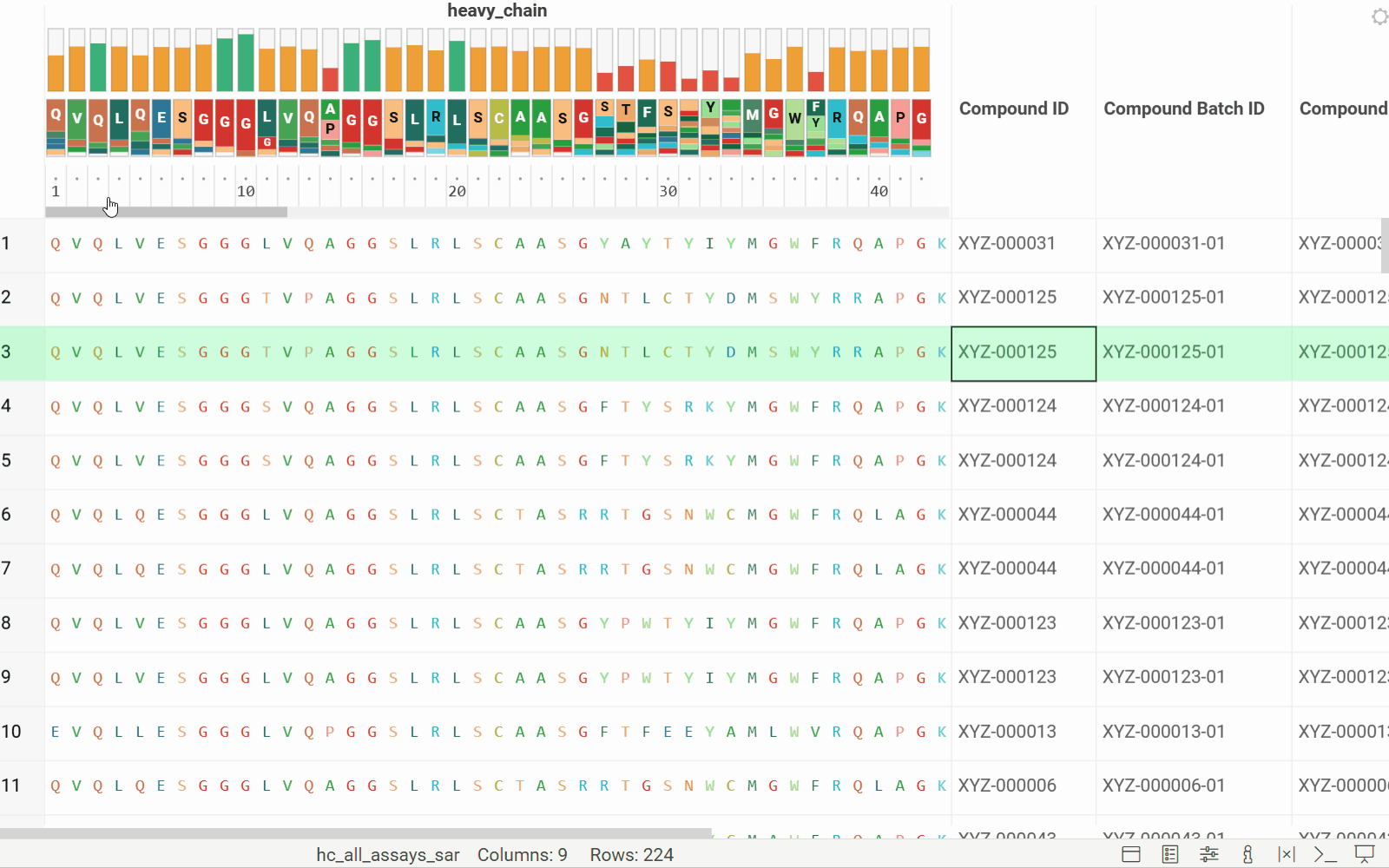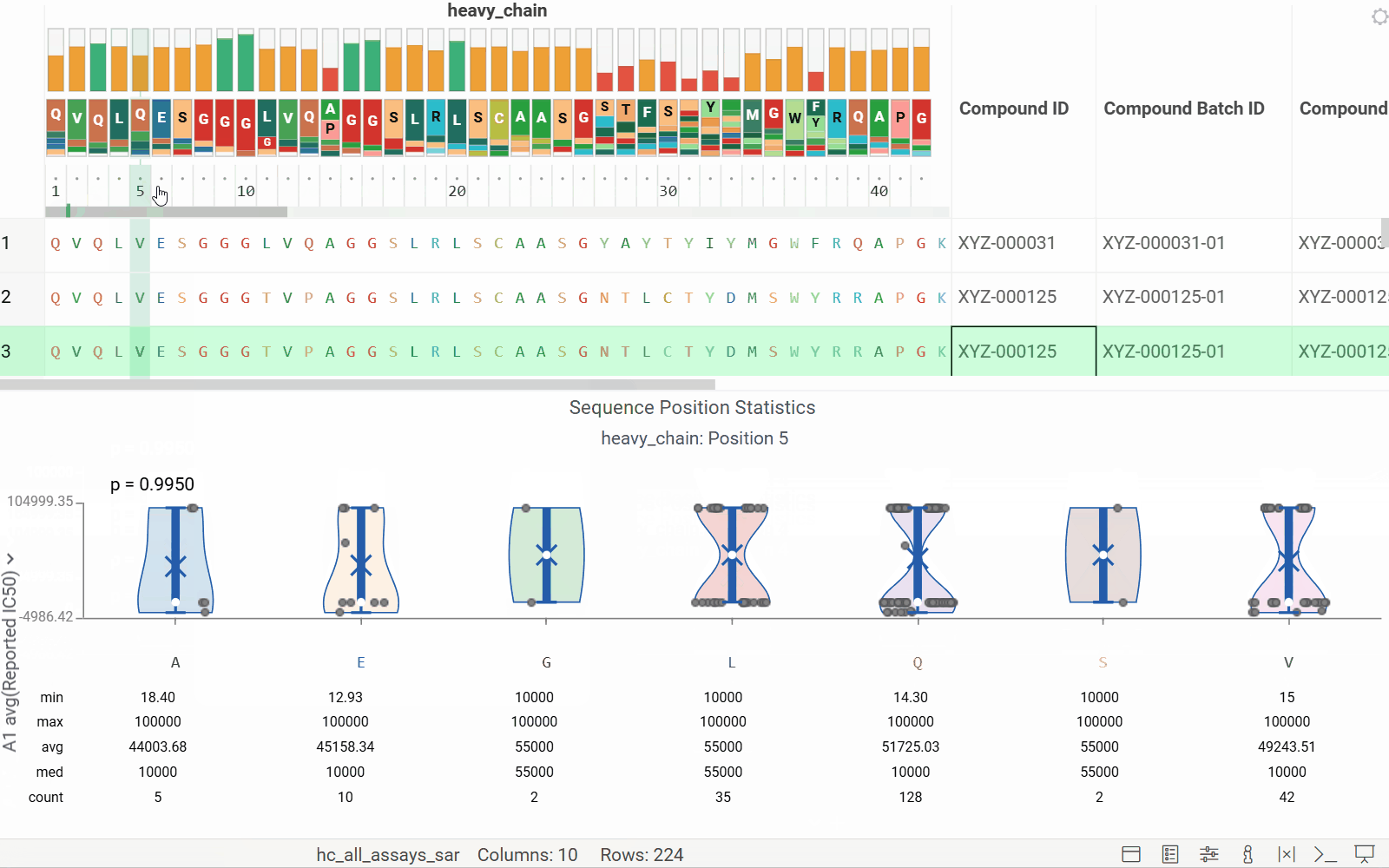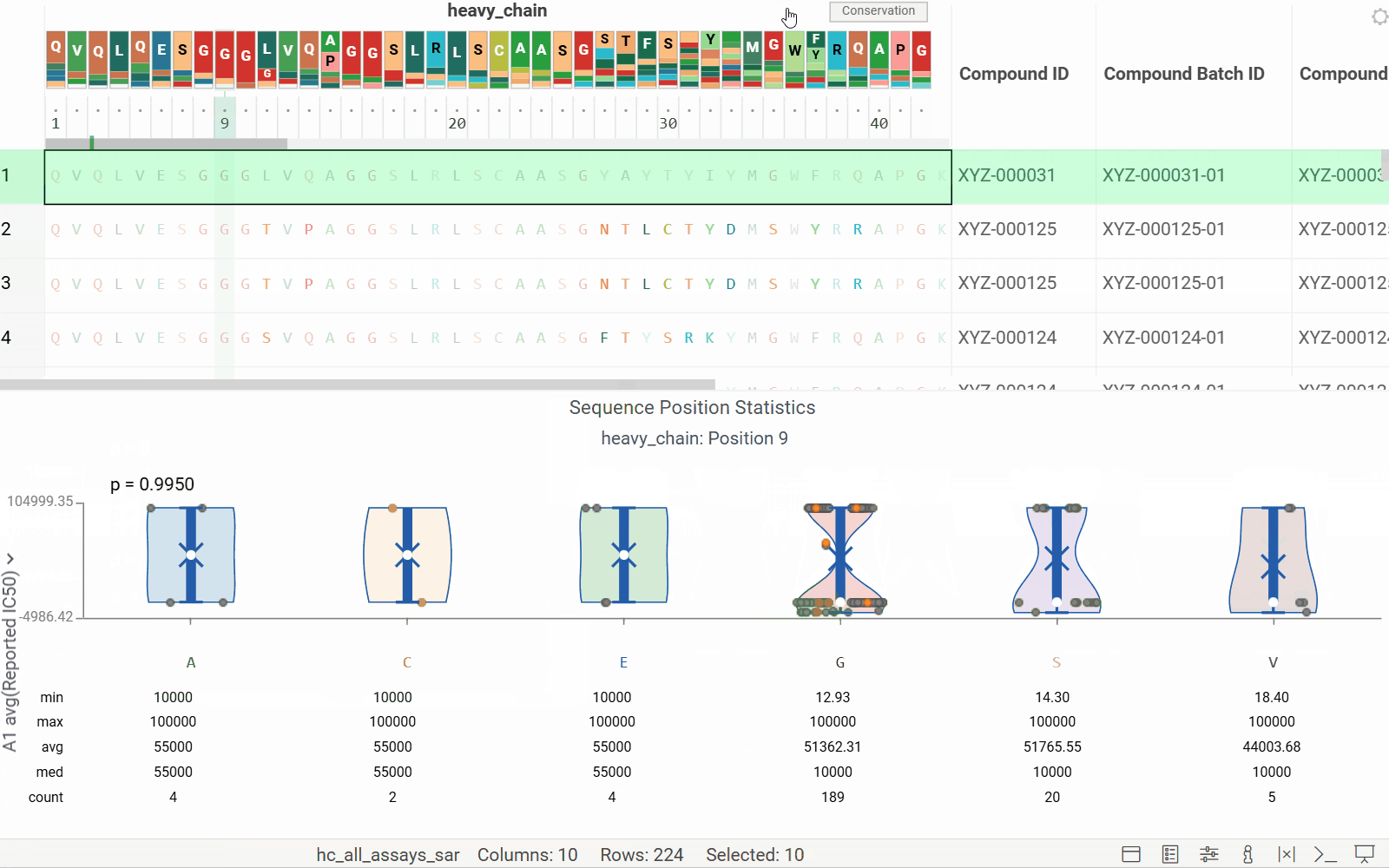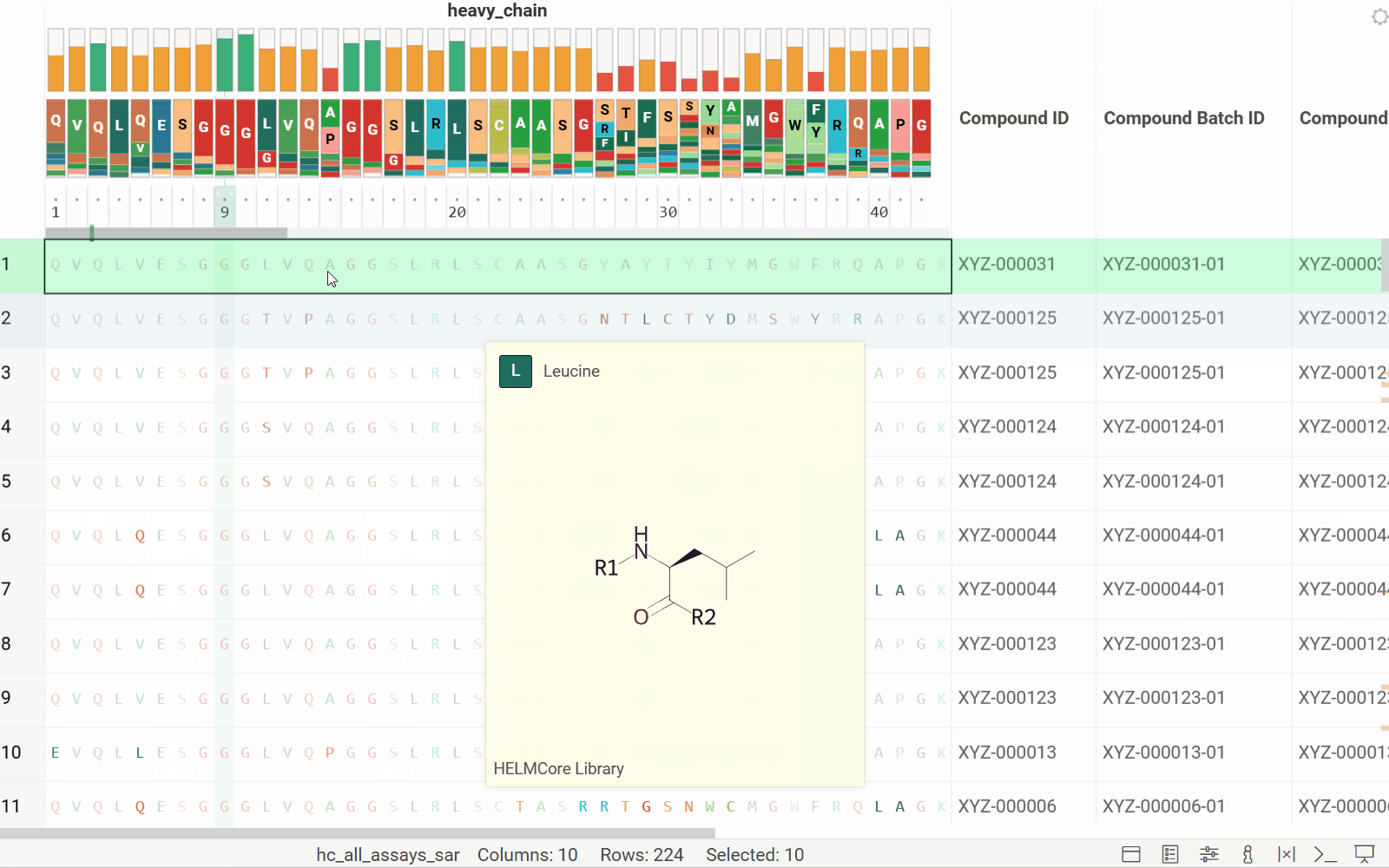
- Added header tracks: Conservation and WebLogo
- Added tooltips for the WebLogo header
- Improved performance of Conservation and WebLogo calculations
Sequence Scrolling and Positioning:
- Added optional scrolling header for short sequences
- Improved detection of the maximum sequence length
- Improved display of the current position
Performance Improvements:
- Added faster methods for retrieving monomers at specified positions
See the new MSA header enhancements in action:
2 Likes
File Editors 1.3.1 (2025-07-30)
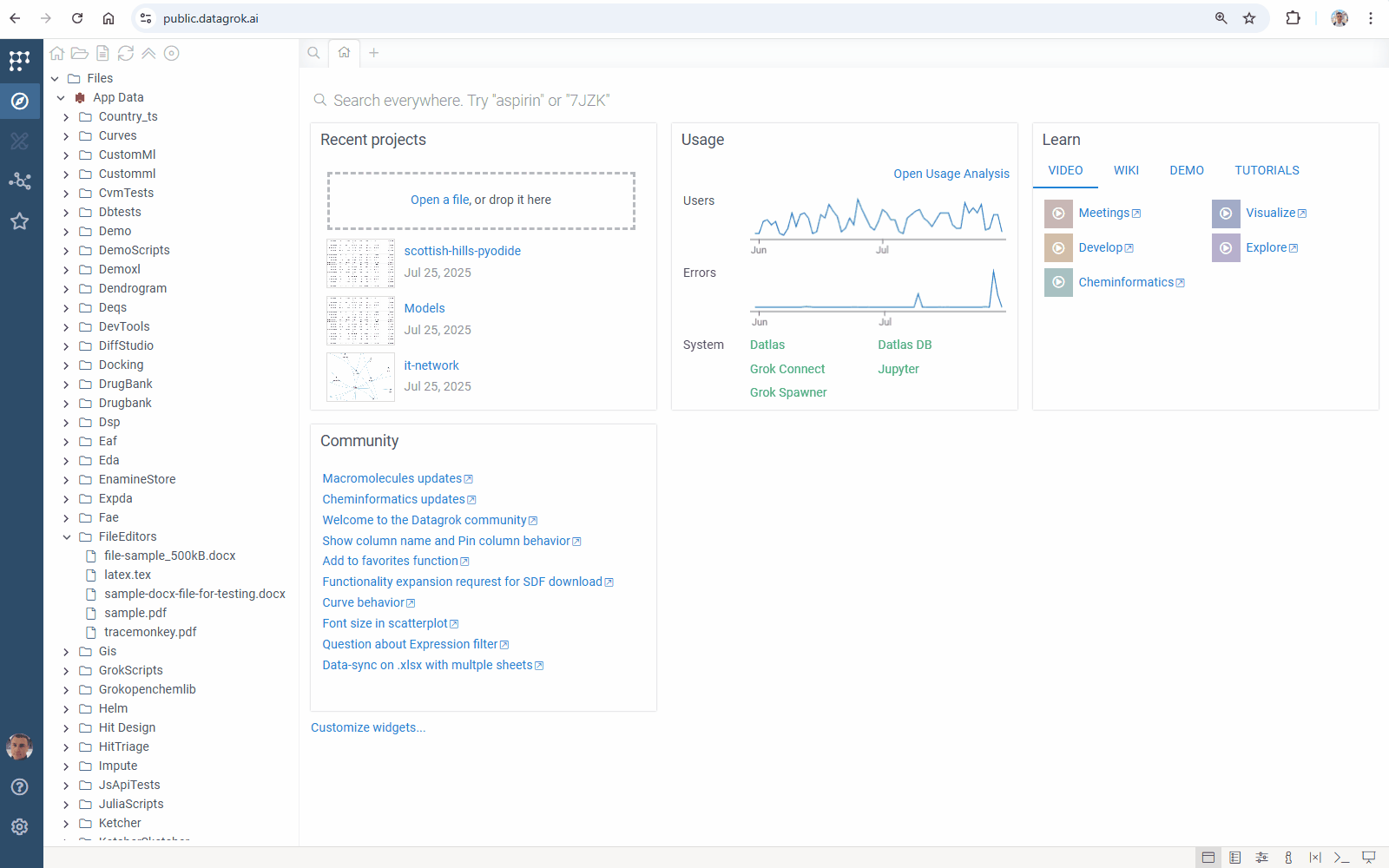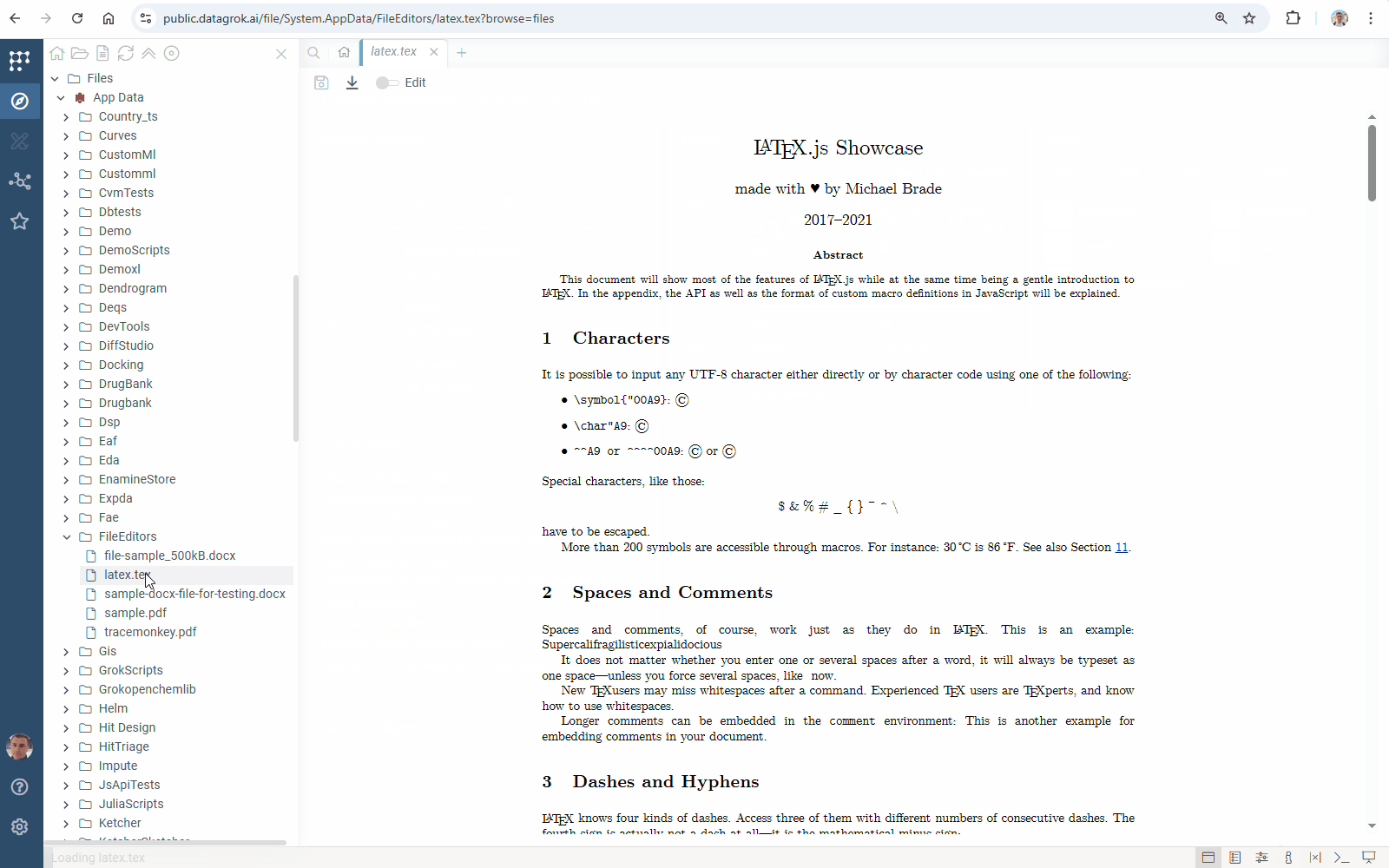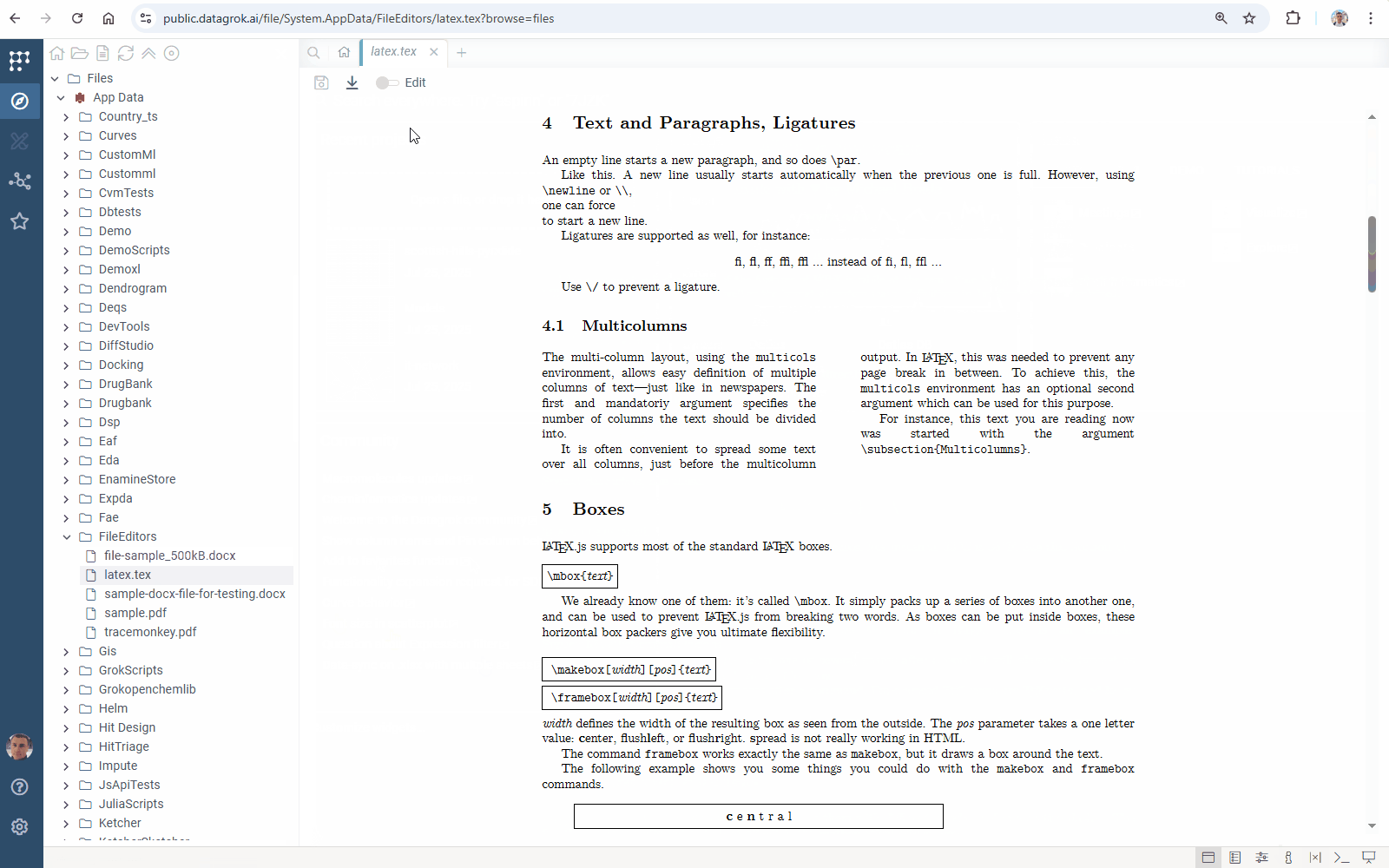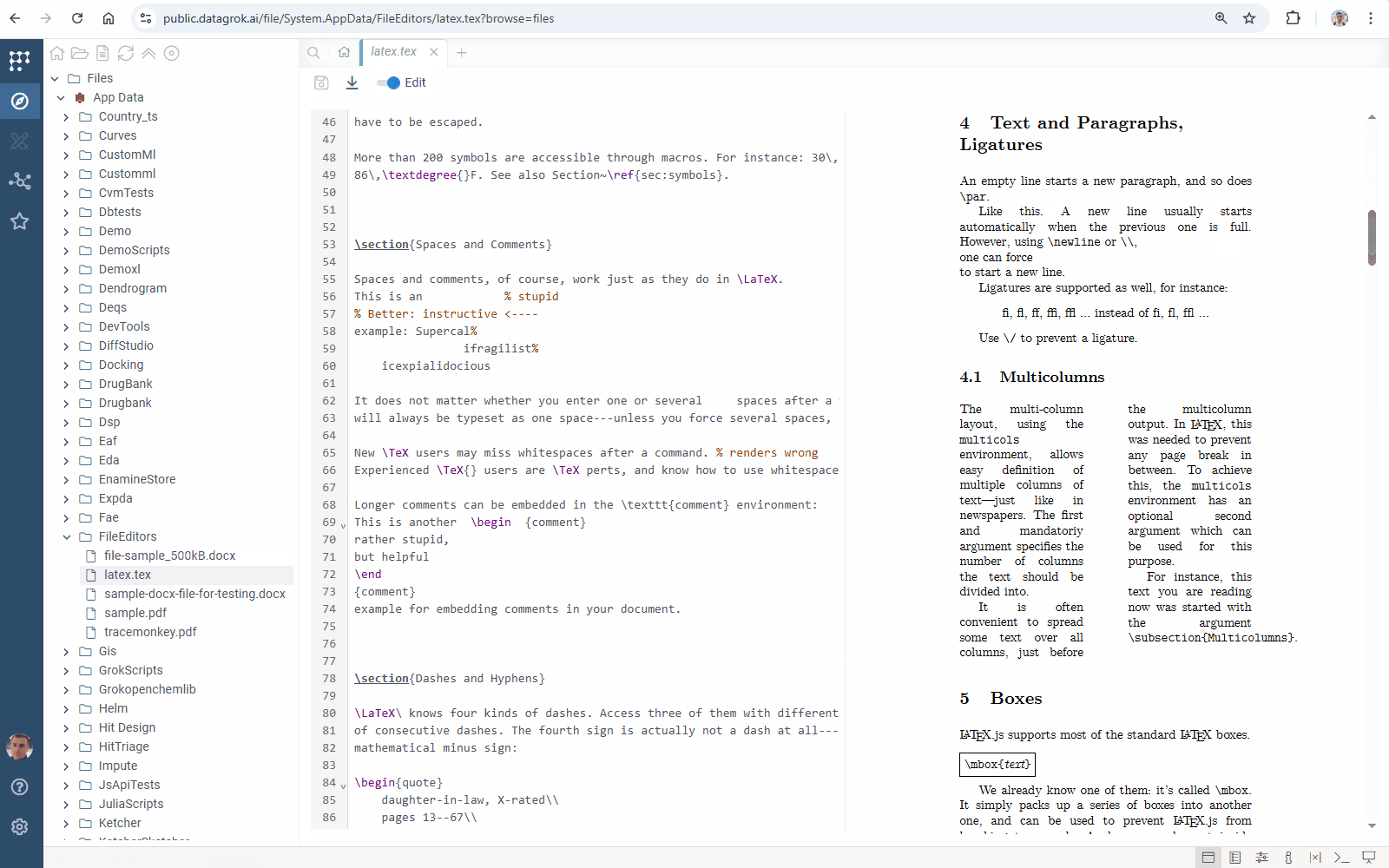
Added TeX format support with full viewing, editing, and sharing capabilities.
3 Likes
Pyodide 1.3.0 (2025-07-25) docs, release notes
Features:
- Support calling
FuncCalls using run_sync and execFuncCall
- Support script dependencies using the
meta.dependencies annotation
- Display script errors in the console with accurate line numbers
1 Like
Curves 1.10.0 (2025-07-29) docs, release notes
Features:
- 4plDoseResponse: Added a function for 4PL dose-response curve fitting
- Data-to-curves: Added full support for the script, including history, data sync, and two-tier support
- Added support for rendering raw PNG images
3 Likes
Chem 1.16.0 (2025-07-28) docs, release notes
Features:
- Mixtures: Added renderer, widgets, and semantic type. For detail, see Cheminformatics updates
- #3426 CPC: Moved to chem, factored into a separate widget file
- #3302: Multiple scaffold trees: Naming/coloring should be saved in projects
- MMP:
- Added hints
- Added collaborative filtering with parent table
- Added the ability to filter fragments/pairs
- View results on PC plot, select activity difference function with scaling, in cell pc plots for activities, show trellis filters in popup, added trellis legend
- Added description to scatter plot for chemical space and activity cliffs
- Similarity/Diversity: Show error in case no molecule column was found
Bug fixes:
- #3408: Scaffold tree: Molecule coloring disappears after closing the tree
- #3346: Export as SDF: Fixed none values are written as numbers ant not as empty strings
- #3399: Do not allow to modify Scaffold colors from Scatterplot or other viewers
3 Likes
Charts 1.6.0 (2025-07-28): docs, release notes
Bug fixes:
- 3420: Tree viewer: Moved
Show mouse over line to Style section on the Context Panel
- 3421: Tree viewer: Different row source behavior when changing
On Click property
- 3397: Tree viewer: Selection and clicking issues
2 Likes
Sequence Translator docs, release notes
The Sequence Translator plugin is a specialized bioinformatics tool that enables translation of oligonucleotide sequences between different representations and provides advanced polymer sequence manipulation capabilities.
Sequence Translator consists of two main components:
- Oligo Toolkit: Translates oligonucleotide sequences between different formats used by various synthesizers (Axolabs, BioSpring, MerMade12, etc.)
- PolyTool: Handles cyclized peptides and polymer sequences with specialized conversion and enumeration capabilities
Main features:
- Sequence translation:
- Translates between multiple oligonucleotide formats
- Bulk sequence conversion with table input support
- Chemical structure generation (SDF, SMILES) from sequences
- PolyTool advanced features:
- Cyclized peptide sequence conversion from custom notation to HELM format
- Sequence enumeration for combinatorial chemistry applications
- Rule-based sequence manipulation with JSON configuration files
- Atomic-level molecular structure conversion
- Integration features:
- Custom cell renderers for cyclized sequences in grid and other viewers
2 Likes
Bio 2.22.12 (2025-09-30) docs, release notes
Features:
- Added matching with the Monomer Library tool
- Improved converter functionality
- Harmonized options in Similarity/Diversity viewers
- Enabled forced detection of macromolecules for arbitrary columns
2 Likes
Chem 1.16.1 - 1.16.5 (2025-09-15) docs, release notes
Features:
- Added persistent layout tag for scaffold alignment
Bug fixes:
- #3464 Double values in SDF export are saved with unnecessary precision
- #3445 Structure filter does not realign when coordinates change but structure does not
- #3433 MMP viewer can’t be added in case with multiple tabs
2 Likes
Sequence Translator 1.9.14 (2025-09-23) docs, release notes
Improved Polytool rules editor, added more examples
2 Likes
Curves 1.10.1 - 1.10.12 (2025-09-16) docs, release notes
- Enabled skipping rendering too small plates
- Data-to-Curves:
- Added well-level additional columns
- Added joining options
- MultiCurveViewer: Added
showOutliers parameter and legendColumnName property
- Added MSR script and updated environment
2 Likes
Power Grid 1.5.5 - 1.7.9 (2025-09-23) docs, release notes
Features:
- Forms viewer: Added substructure highlighting support
- Smart form: Enabled display of only the first ten columns by default
Bug fixes
- Smart form:
- Fixed issue with adding categorical columns from the Context Panel
- Fixed incorrect rendering on column resize
2 Likes
Power Pack 1.4.0 - 1.7.7 (2025-10-02) docs, release notes
Features:
- Implemented the Activity dashboard widget
- Complex calculated columns: Enabled persisting with the layout
Bug fixes:
- #2449: Add new column: Using non-empty rows in preview
- #3306: Calculated columns: types validation error
- #3389: Formula line preview for scatterplot used in trellis plot does not reflect axes configured in scatterplot
- #3269: Projects: XLSX files with multiple sheets: data sync is only provided for the first sheet
- #3392: Scatterplot: Formula lines: axis in dialog should be the same as on the viewer
2 Likes
Bio 2.23.0 - 2.25.1 (2025-10-30) docs, release notes
Breaking changes:
- Reworked monomer library management: transitioned to library providers instead of single file-based monomer libraries and enabled support for multiple providers. Update your code accordingly
Features:
- HELM support: Added CHEMS /SMILES and multi-polymer type support
- Full BILN support: Added rendering and conversion for SMILES/CHEMS, HELM, separator, FASTA, and molecular forms; parsing; monomer library handling
- Improved monomer library loading speed
Bug fixes:
- Fixed dialogs for adding/removing monomer libraries
- Fixed HELM converter to support multi-type polymers
2 Likes
Chem 1.16.6 (2025-10-16) docs, release notes
Features:
- Improved Mixtures widget in the Context Panel to display component names where available
- Enabled calculation of Context Panel Descriptors via Docker
- Add new column dialog: Added ability to run Descriptors, Properties, Toxicity Risks, and Structural Alerts
- Scaffold tree: Added explanation tooltip why the tree can’t be generated
Bug fixes:
- #3497: Chem: MPO: Update to work correctly with data sync
- Similarity Search viewer: incorrect coloring after sorting a dataset
- Fixed distance functions
2 Likes
Power Grid 1.7.10 (2025-10-21) docs, release notes
Fixed:
- Forms viewer: incorrect row coloring when scrolling grid with pinned rows
2 Likes
HELM 2.11.0 - 2.13.1 (2025-10-31) docs, release notes
Features:
- Monomer library management: transitioned to library providers instead of single file-based monomer libraries and enabled support for multiple providers.
- Added CHEMS and SMILES support
- Helm Web Editor: Added multi-polymer type support
Fixed:
- Compatibility with platform version 1.25
2 Likes
Compute 1.45.5 (2025-10-30) docs
2 Likes
Peptides 1.25.0-1.27.2 ( 2025-11-12) docs, release notes
Features:
- Introduced the sequence mutation cliffs viewer with support for current row tracking
- Enabled full filter reactivity for all panels, tooltips, aggregations, and viewers
- Reworked mutation cliffs and tooltips, showing information about sequence pairs and their statistics
- Added sum columns for the sequence variability map viewer
- Removed target column for peptides analysis
Bug fixes:
- Corrected aggregate calculations, statistics, and mean difference comparisons
- Fixed premature shutdown of MCL viewer on loading
- Fixed distribution tooltips not showing on WebLogo hover
2 Likes

